Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Mass Spectrometry
Resolving power for the people: Ion mobility-mass spec expands its offerings
New instruments come online with improved resolution, broadening applications for the technique
by Celia Henry Arnaud
May 22, 2020
| A version of this story appeared in
Volume 98, Issue 20

When ion mobility spectrometry (IMS) and mass spectrometry (MS) were first combined in a commercial instrument more than a decade ago, users had limited options. If they wanted the analytical power this combination provides, they could either purchase that single instrument, or they could build their own. Today, more than 500 IMS-MS instruments are in laboratories around the world, according to estimates by Strategic Directions International, which tracks the analytical instrument market. The ion mobility units in those instruments come in four configurations rather than one, with a fifth on the way (see page 18).
The explosion in IMS-MS instruments is in response to a demand for more sophisticated chemical analysis. Adding IMS to mass spec instruments enables users to study more complex mixtures than ever before. It provides information about molecules that mass spec alone can’t provide.
MS separates ions on the basis of the mass and charge of ions in a sample, and adding IMS allows users to separate ions on the basis of size and shape too. These extra degrees of separation mean that molecules in a sample can be more thoroughly characterized, leading to fewer false positives in ion identification.
With more types of IMS-MS instruments to choose from, scientists also have more options in the resolution of their measurements. The many instruments available today come with a range of resolving powers, which is a measure of the machine’s ability to distinguish between two nearly identical ions. The higher the resolving power, the better the separation. So for instance, the first two versions of IMS to be coupled with MS—drift tubes and traveling waves—typically achieve resolving powers of 50–60 and 30–40, respectively. Newer types of IMS instruments can achieve even higher resolving powers. Trapped ion mobility spectrometry (TIMS) achieves up to about 400–600 resolving power depending on how fast the electric field is ramped, and cyclic IMS achieves a resolving power of about 750 by passing ions some 100 times around its 1 m long track. Structures for lossless ion manipulations (SLIM) devices, the newest of the bunch, are built on printed circuit boards, which lengthen the path ions take through the compact devices. This type of device could hypothetically achieve a resolving power of 2,000 for ions passing 100 times over its 13 m path length.
The ion mobility toolbox
C&EN takes a spin through the catalog of instruments now available to enhance the molecular resolving power of mass spectrometry with ion mobility.
Adding an ion mobility device to a mass spectrometer can improve the machine’s resolving power, meaning that it can distinguish more clearly among the ions it’s analyzing. Some of the first ion mobility units to be added to mass spec instruments were drift tubes, which separate ions based on their collision with buffer gases like helium inside the tube. In these devices, the larger the ion, the larger its so-called collision cross section, and the longer it stays in the tube before eluting. These illustrations highlight the many ion mobility devices offered by companies today—from simple drift tubes to cyclic racetrack configurations.
Ion resolving power: 30–40 | First commercialized by: Waters in 2006
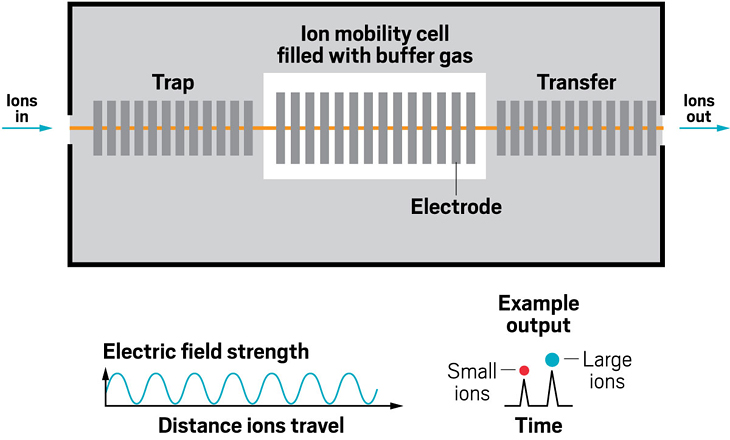
Ion resolving power: 50–60 | First commercialized by: Agilent Technologies in 2014
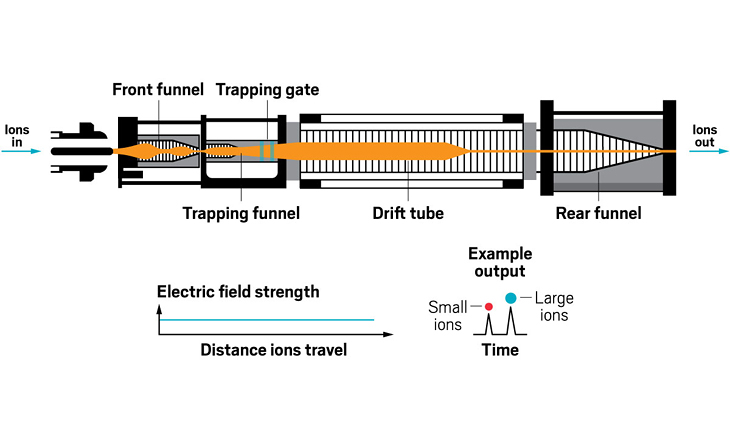
Ion resolving power: Up to 400, with slow ramping | First commercialized by: Bruker Daltonics in 2016
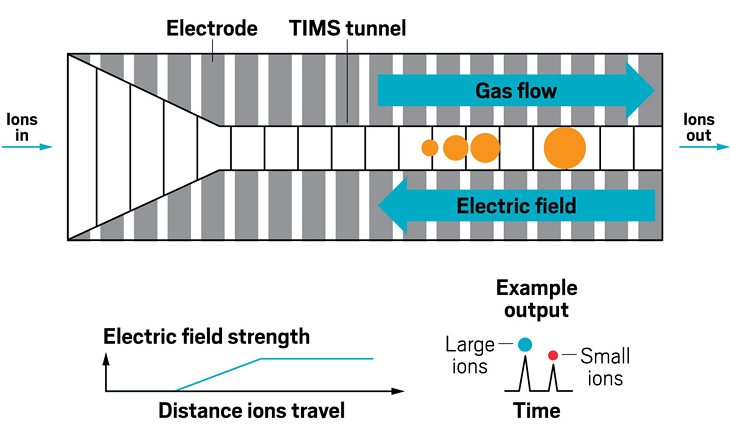
Ion resolving power: ~750, by sending ions around the cycle 100 times | First commercialized by: Waters in 2019
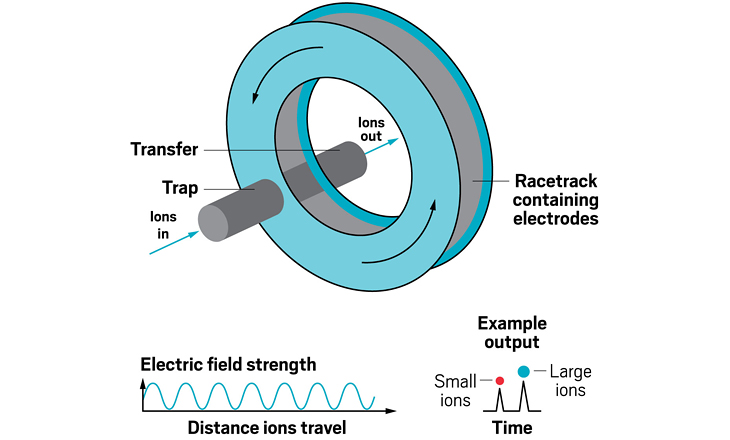
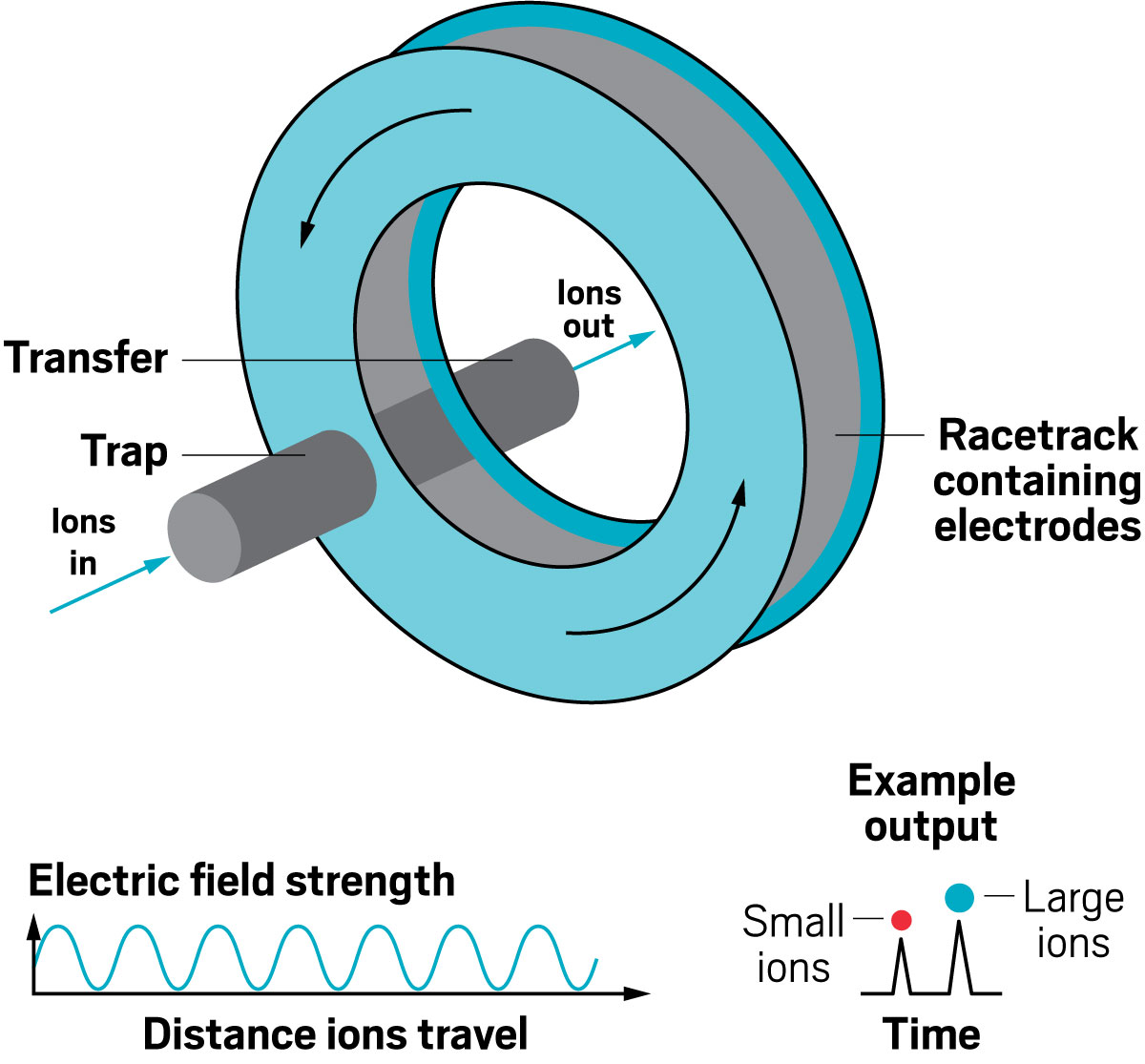
Ion resolving power: 200–300 for a 13 m path length; up to 2,000 with multipass devices | To be commercialized by: MOBILion Systems, with a projected launch in June 2021
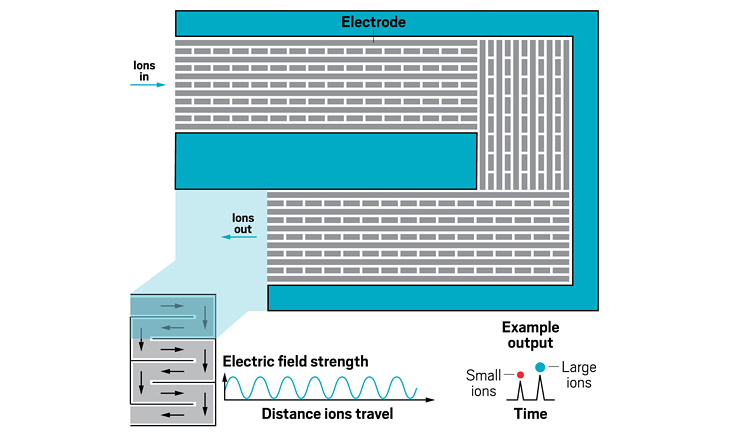
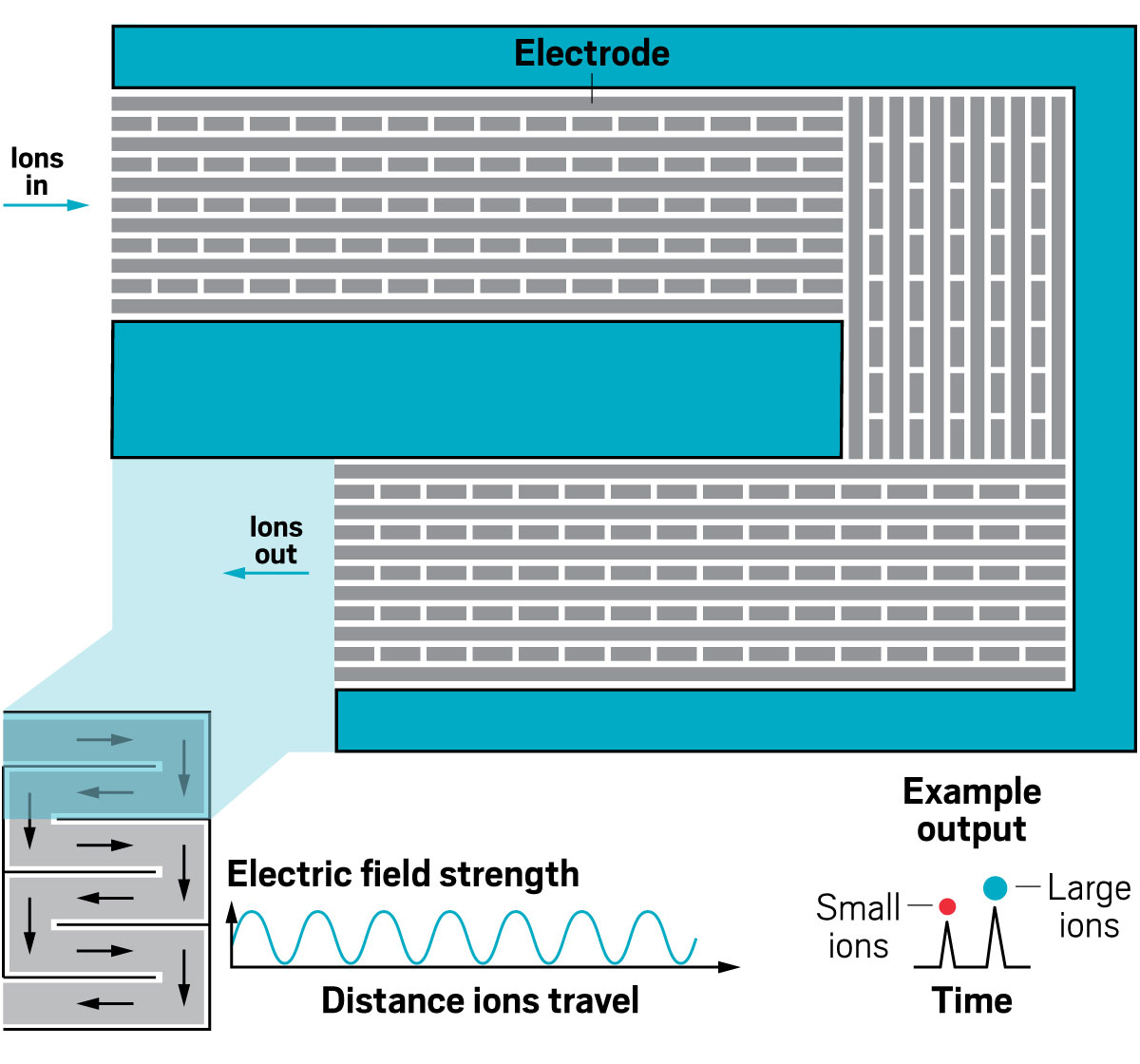
The high resolving powers of the various instruments open up new abilities for analyzing molecules of all sizes. For large molecules, IMS-MS is providing information about the landscape of configurations a protein may take. For small molecules, IMS-MS is being used to separate mixtures of closely related species like isotopomers.
Isotopomers are compounds that have the same molecular formula and contain the same elemental isotopes. The only difference between them is the location of the isotopes within the compounds—which atomic positions they reside on. For instance, two otherwise identical organic compounds could have a deuterium on a terminal carbon or an internal carbon.
Richard D. Smith and coworkers at the Pacific Northwest National Laboratory use SLIM devices to tease apart isotopomers as part of their efforts to better understand how IMS separations work. The team confirmed that SLIM could distinguish between isotopomers.
“At first blush, you would say these things should have the same mobility. They have the same shape, same charge, same mass,” Smith says. But the difference in the location of the isotopes is enough to change the center of mass and the moment of inertia of the molecules. Those changes lead to slight differences in mobility that can be seen with high enough resolving power. And such high resolving power has many practical applications, including separations of structurally similar lipids, glycans, and many isomers that have previously not yielded to separation by IMS.
IMS is finding applications in forensics as well. Francisco A. Fernández-Lima, a chemistry professor at Florida International University, uses TIMS to help his forensics colleagues analyze structurally similar drugs such as cannabinoids and synthetic opioids. Many of these drugs are positional isomers, which means they have the same scaffold with a functional group appearing at different positions across that scaffold. The drugs’ potency depends on which isomer form they take on. “It’s not a mass problem,” Fernández-Lima says. “It’s a structural problem.”

Fernández-Lima also uses IMS-MS to study complex environmental samples such as crude oil and dissolved organic matter. Typically the readout from IMS or MS measurements gives users information about the potential identity of ions in their samples—it gives them chemical formulas but not exact configurations. In Fernández-Lima’s environmental experiments, he combines TIMS with a type of mass spectrometry called ultra-high-resolution Fourier transform ion cyclotron resonance MS that provides such high mass resolving power that he can determine unique molecular formulas. “It narrows a lot more the potential candidates you actually have for that chemical formula,” he says.
Another environmental application for IMS-MS is the analysis of per- and polyfluoroalkyl substances (PFAS). These pollutants have been in the news because of their reputation as “forever chemicals.” Long-term emissions from a Chemours plant in Fayetteville, North Carolina, contaminated the Cape Fear River with PFAS.
IMS expert Erin S. Baker and PFAS expert Detlef R. U. Knappe, both at North Carolina State University, added drift tube IMS to an existing workflow for analyzing PFAS (Anal. Chem. 2020, DOI: 10.1021/acs.analchem.9b05364). With the IMS, they were able to separate branched and linear PFAS with identical masses but different shapes. Such analyses that identify specific pollutants in local water samples could help inform public policy.
Baker and Knappe can also separate PFAS into classes distinguished by their head groups. “This allows you to distinguish known PFAS and identify new PFAS in untargeted analyses of biofluid, tissue, and environmental samples,” Baker says.
Because PFAS impair lipid function, Baker plans to combine IMS-based PFAS analysis with lipidomics. Lipids, another complicated mixture with many structural isomers, can benefit from mobility separations. She wants to see how PFAS exposure affects the various kinds of lipids in cells.
IMS provides extra information and separation that makes screening applications easier, such as identifying the products of enzyme reactions occurring inside cells. “If we screen based on mass and mobility, we have much better target hit rates for our compounds,” says Perdita E. Barran, a chemistry professor at the University of Manchester. In monitoring enzyme reactions, mobility measurements help her team easily distinguish between closely related substrate and product ions.
Barran would like to see clinical instruments that combine high-resolution IMS with low-resolution mass spectrometry. “Ion mobility is cheap,” she says. “Mass spectrometry is expensive” because it requires costly vacuum pumps. Combining the techniques in this way would enable labs that couldn’t normally afford a high-resolution MS instrument to achieve reasonably good results.
Advertisement
John A. McLean of Vanderbilt University is combining IMS-MS with desorption electrospray ionization imaging to rapidly screen metabolites in genetically altered cyanobacteria. “The genetic alterations can be made really, really fast,” McLean says. But “knowing the consequences of those changes is the real bottleneck.” Researchers want to use genetically modified organisms like cyanobacteria as chemical factories, programming them to synthesize desired molecules. When testing their designer organisms, the scientists need to ensure the modifications they’ve put in place are performing as intended, and they do that by monitoring the bacteria’s metabolites.
McLean and colleagues use CRISPR gene editing to alter cyanobacteria’s biological clock machinery and tweak the microbes’ metabolic pathways. They use imaging IMS-MS analysis to determine which metabolites the organisms produce.
The applications don’t stop with small molecules, however. IMS-MS is being used for multiple protein-related applications, from proteomics to structural biology.
For Matthias Mann’s groups at the Max Planck Institute of Biochemistry in Munich and the University of Copenhagen, adding TIMS to their existing proteomics workflow has allowed them to achieve better sensitivity, which in turn increases the number of peptides they can detect and quantify. They developed a method called parallel accumulation-serial fragmentation (PASEF) that allows them to accumulate multiple precursor ions in the TIMS device and then release them sequentially into the mass spectrometer for tandem MS analysis (Mol. Cell. Proteomics 2018, DOI: 10.1074/mcp.TIR118.000900). “You get a huge signal-to-noise improvement,” Mann says.
In addition to using IMS to confirm the identity of regular peptides he’s studying, which reduces false discovery rates in proteomics experiments, Mann says the technique can help identify cross-linked peptides. These peptides have a larger collision cross section, a parameter measured by IMS that’s related to an ion’s shape and conformation, than other peptides.
“They have a bigger cross section, so they stick out in the graph,” he says, referring to where the cross-linked peptides show up in the IMS-MS instrument’s output.
IMS-MS has also long been a tool for structural biology of proteins and protein complexes.
Christian Bleiholder of Florida State University combines two TIMS cells for top-down analysis of intact proteins, instead of digesting them into fragments first, a common practice in protein mass spec analysis. Bleiholder selects a species or protein conformation using the first TIMS cell. Then he uses a high electric field to break apart the ion and separate the fragments with the second TIMS cell.
“We can do mobility-selective top-down sequencing—the ability to get structure and sequence at the same time,” Bleiholder says. He’s particularly interested in using the approach to analyze proteins with posttranslational modifications such as sugars. He’s still working on developing the method, but he eventually wants to apply it to disease-related proteins such as glycoproteins from HIV or the novel coronavirus, SARS-CoV-2.

In another protein-related application, researchers use IMS-MS to monitor collision-induced unfolding (CIU) of antibodies. Brandon Ruotolo and coworkers at the University of Michigan measure changes in the collision cross section of antibodies as a function of the energy. In this way, they get a fingerprint of the intermediate states an antibody goes through on its way from being fully folded to fully unfolded. Ruotolo is developing the method as a way for biotherapeutic manufacturers to ensure that the antibodies they’re producing are in a specific configuration.
With high-resolution mobility measurements, he can get an even more detailed CIU fingerprint. He selects conformations that were at low levels in the original fingerprint, enriches them using IMS, and measures the CIU of that subset of ions. “You tend to get new features that are not well represented in the first fingerprint,” Ruotolo says. In that way, a fingerprint with four or five features can be expanded to one with 10–20 features.
Sectors like the pharmaceutical industry may also soon benefit from improvements to IMS-MS. MOBILion Systems is collaborating with Agilent Technologies to launch the first commercial SLIM instrument. With a focus on improving the characterization of biopharmaceuticals, they’re looking to use the high resolving power of SLIM to reduce the need for liquid chromatography (LC) in applications such as glycan analysis, peptide mapping, and intact protein analysis, says Melissa Sherman, MOBILion CEO.
“For complex samples, we may not eliminate LC altogether, but we may use it in a different way,” Sherman says. “We’re using it to filter and spread the sample out to optimize the ionization.”
As the popularity of IMS-MS continues to surge and more instruments come online, the user community is coming together to improve measurements for everyone. One piece of information that IMS-MS can provide is the collision cross section of various conformational states of molecules. Typically, drift tube devices measure absolute cross sections and can be used to calibrate other instruments. But most labs don’t have multiple types of IMS instruments. To make such calibration easier for labs that don’t have a drift tube instrument of their own, McLean started the Unified Collision Cross Section Compendium as a way to crowdsource IMS data that people can use to calibrate other types of systems (Chem. Sci. 2018, DOI: 10.1039/c8sc04396e). Every value in the compendium goes through a rigorous validation process.
“To help us validate entries in the CCS Compendium, we have also compiled a literature database of every cross section published over the past 100 years,” McLean says (Anal. Chem. 2016, DOI: 10.1021/acs.analchem.6b04905). The compendium itself has collision cross section data for about 4,000 compounds. It’s available for download so that researchers can incorporate it into their bioinformatics workflows. It has also been incorporated into databases from the US National Institute of Standards and Technology and other organizations, McLean says.
These applications are just some of the many to which IMS-MS is being applied. Who knows how many more uses researchers will come up with as the technology continues to improve.
CORRECTION
This article was updated on May 29, 2020, to correct the description of the Cape Fear River PFAS pollution. It was the result of long-term emissions, not a 2017 spill.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter