Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Structural Biology
Covid-19
How structural biologists revealed the new coronavirus’s structure so quickly
Scientists detail the steps they took to determine the structures of SARS-CoV-2’s proteins and the next steps toward COVID-19 treatments
by Laura Howes
May 2, 2020
| A version of this story appeared in
Volume 98, Issue 17

When the government in Wuhan, China, confirmed on Dec. 31, 2019, that authorities there were treating dozens of cases of pneumonia of unknown origin, researchers who remembered the severe acute respiratory syndrome (SARS) outbreak of 2003 were uneasy. Was this another coronavirus, like the one that spurred that incident? Slowly, concern spread throughout the scientific community, and labs that had already been studying coronaviruses began to get primed.
Jason S. McLellan, a structural biologist at the University of Texas at Austin, remembers getting a call while on a skiing vacation with his family. On the other end of the line was Barney S. Graham, deputy director of the US National Institutes of Health’s Vaccine Research Center. The pair had worked together in the past, and Graham told McLellan: “It looks like it’s coronavirus. Are you ready to get to work on it?” McLellan says he messaged his team via the mobile service WhatsApp and told them to get ready: “We’re going to race as soon as we get the sequence.”
Support nonprofit science journalism
C&EN has made this story and all of its coverage of the coronavirus epidemic freely available during the outbreak to keep the public informed. To support us:
Donate Join Subscribe
The sequence McLellan was referring to was the genomic sequence of the new coronavirus that Graham told him about. With the virus’s genome in hand, McLellan and his team would be able to synthesize its most vital proteins and then determine their structures, an important first step in finding therapeutics to fight a pathogen. By early January, the wait was over. A team led by researchers at Fudan University had published the genome, sharing it publicly online so that labs around the world could leap into action before the related paper was published (Nature 2020, DOI: 10.1038/s41586-020-2008-3).
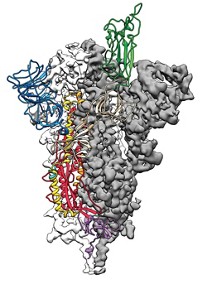
And McLellan’s team did indeed race. The research group was one of the first to publish a cryo-electron microscopy structure of one of the new coronavirus’s proteins. The scientists determined the configuration of the virus’s spike protein, a biomolecule that decorates the outer shell of the virus and enables it to fuse with and enter human cells to cause infection (Science 2020, DOI: 10.1126/science.abb2507).
Unlike human genetic information, which is encoded in double-stranded DNA, the new coronavirus, like all coronaviruses, stores its genetic information in a single strand of RNA. The human genome contains around 3 billion base pairs tightly packaged inside the nucleus of each of our cells. In contrast, the new virus’s RNA genome has fewer than 30,000 bases.
That shorter sequence, scientists would learn, codes for the 29 proteins that make up the virus, now dubbed SARS-CoV-2. These biomolecules protect the pathogen, help it attach to host cells, and enable it to replicate.
Uncovering the structures of proteins like these helps scientists develop small molecules, antibodies, and other therapeutics that can disrupt the proteins’ function. Although the process of determining a protein structure has gotten easier over the years because of advances in technology and know-how, the speed at which teams uncovered SARS-CoV-2’s protein structures was unprecedented.
“I think this is really exceptional,” says Sarah J. Butcher, a structural biologist at the University of Helsinki. “It was 5 weeks after the first cases of COVID-19 began to appear that the first structures were deposited in the PDB,” she says.
The PDB is the Protein Data Bank, an internationally managed database for the 3-D structural data of large biological molecules. According to statistics from the PDB, by the end of March, the service has already received over 100 structures related to the new coronavirus. And submissions keep coming.
The importance of experience
The research groups that won the race to solve the first protein structures of SARS-CoV-2 were the ones that had worked on past coronaviruses, says David R. Armstrong, a member of the PDB’s European team, which is based at the European Bioinformatics Institute in the UK. Those groups include McLellan’s in Texas, Rolf Hilgenfeld’s at the University of Lübeck, and Zihe Rao’s at Tsinghua University. Hilgenfeld and colleagues elucidated an early structure of the virus’s main protease, an enzyme that helps SARS-CoV-2 build its proteins (Science 2020, DOI: 10.1126/science.abb3405), and Rao and team determined structures of both the virus’s protease (Nature 2020, DOI: 10.1038/s41586-020-2223-y) and its polymerase (Science 2020, DOI: 10.1126/science.abb7498), which helps SARS-CoV-2 make copies of its RNA genome.
Those labs had years of shared experience working on the protein structures for related viruses, like the coronaviruses that cause SARS and Middle East respiratory syndrome (MERS), a disease first reported in Saudi Arabia in 2012. Because these viruses are similar to SARS-CoV-2, the scientists could look at the RNA sequence of the new coronavirus and immediately find the sections coding for the proteins they were interested in. The shapes of the proteins looked similar too.
Numerous experts have told C&EN that the biggest challenge for structural biologists today is to get well-behaved protein samples that are stable enough to undergo structural characterization. That SARS-CoV-2 was similar to the viruses that caused SARS and MERS outbreaks was a boon: labs already knew how to make and stabilize these proteins. Researchers could order pieces of DNA that code for SARS-CoV-2’s proteins and be confident that they would work in processes already established in the lab. Once they received the DNA, the scientists could insert it into cells grown under the right conditions, and those cells could churn out copies of the desired protein.
When recounting the early days of the outbreak of COVID-19, the disease caused by SARS-CoV-2, McLellan pulls up a timeline on his computer: “Jan. 30 was a big day,” he says. That was the day graduate student Daniel Wrapp harvested the team’s spike protein and purified it. The team had earlier inserted a DNA sequence—in the form of a circular construct called a plasmid—into some mammalian cells, reprogramming their machinery to synthesize the spike protein. “Later that night,” McLellan says, Wrapp “started freezing grids, and we began the initial rounds of data collection.”
The grids are part of a cryo-electron microscope—one of the two main tools structural biologists use to determine protein structures.
“Oh yeah,” Cynthia Wolberger, a structural biology expert at Johns Hopkins University School of Medicine, says, laughing when she hears that all those steps happened in 1 day. “Once you have a purified, well-behaved protein, you can just throw it right on the grid.”
“I used to say that in structural biology, one-third of the work was sample preparation, one-third of the work was microscopy, and one-third of the work was image processing,” the University of Helsinki’s Butcher says. Because of myriad technology advances, today “it can be about 80% sample preparation, 10% imaging, and 10% image processing,” she says. “Both image processing and imaging have improved tremendously.”

Technological advances
In 1912, William Lawrence Bragg showed that X-rays could be used to reveal the atomic structures of crystalline salts. By the 1950s, that same technology was starting to probe the structures of DNA and proteins. The field of structural biology was born. Since then, fuzzy photographic plates of X-ray diffraction patterns and blobby, low-resolution models of biological molecules have given way to near-atomic-resolution images.
For years, the workhorse of structural biology has been X-ray crystallography. If researchers can make enough protein and get that protein to crystallize in a highly ordered lattice, the technique can reveal a 3-D structure of the protein. The crystal is rotated through an X-ray beam, and the resulting diffraction patterns are transformed into a map of electron density that reveals the protein’s structural secrets. Crystallographers combine that map with the amino acid sequence to build a model of how the protein folds into sheets and helices.
Today, a lot of the work of protein crystallography is automated. Liquid-handling robots can perform miniature crystallization experiments to find the right solution conditions for growing protein crystals. Synchrotron X-ray sources equipped with microfocused beamlines and cryocooling instruments protect protein crystals from radiation damage, enabling their analysis. Data can now be gathered rapidly from crystals that would have been discarded as too fragile or small 10 years ago. Today, solid-state hybrid pixel detectors on X-ray instruments can capture more than 100 diffraction pattern images per second. “If the crystals are good,” says Manfred Weiss, head of macromolecular crystallography at the Helmholtz-Zentrum Berlin für Materialien und Energie (HZB), “you can collect a good data set within 10–15 min or so.” Fifteen years ago, the same data collection could have taken a few hours.
Weiss has spent years optimizing the protein crystallography work at the BESSY II synchrotron in Berlin. At the end of January, he took a phone call from the University of Lübeck’s Hilgenfeld, the leader of one of the teams that had long been studying coronavirus proteins. Weiss recalls Hilgenfeld telling him, “We have crystals of this coronavirus protease, and we need beam time really urgently.” Recognizing the importance of the request, Weiss complied: “I basically identified a free slot and gave them a beamline 3 days later.” Requests for beamline time at synchrotrons are usually made in writing weeks or even months in advance.
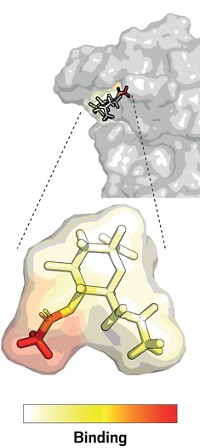
Thankfully, the enzyme that Hilgenfeld’s team was trying to get a structure of had a similar configuration to that of the protease from the virus involved in the MERS outbreak in 2012–13, so the scientists could take some shortcuts to speed up the structure determination process. Linlin Zhang, a postdoc from Hilgenfeld’s lab, took the samples up to Berlin on the Saturday Weiss had assigned the team beam time. By mid-February, the researchers had a crystal structure of the new virus’s protease. That map of the protein’s curves and crevices meant the team could then optimize an existing α-ketoamide inhibitor as a potential drug candidate.
Another technology that’s been key during the COVID-19 crisis is crystallographic screening. In the past 8 years or so, HZB’s Weiss says, the technology has advanced to the point where scientists can quickly cocrystallize a selection of small molecules with protein samples to learn whether one of the compounds binds to a target protein—a step toward designing a therapeutic. Industry has been using screening techniques for a long time, Weiss says, although crystallography has not been a primary screening technique for companies. More than just a yes or no answer as to whether a compound binds, he explains, crystallography can give information about how the molecule binds.
Weiss is screening compounds against the SARS-CoV-2 protease at the synchrotron in Berlin. At the Diamond Light Source, a national synchrotron facility in Oxfordshire, England, a group led by Frank von Delft is also screening compounds against the new protease.

When Tsinghua University’s Rao released a crystal structure of the main SARS-CoV-2 protease bound to an inhibitor in January, his facility was already beginning a scheduled shutdown. Rao passed along what he had learned to the team at the Diamond Light Source so the work could continue. Von Delft’s team focused its efforts and, in 2 weeks, completed the first screen of inhibitors for the SARS-CoV-2 protease. The result was 66 small molecules capable of binding covalently and noncovalently to the active site of the protein. The researchers submitted their data to the PDB and quickly published the results on their website rather than writing a journal article. The achievement was, von Delft says, the result of a lot of “very smart people—very sharp, focused—working together as a collective.”
Yet X-ray crystallography doesn’t work for every protein. Some can’t be crystallized. For these, scientists often turn to cryo-electron microscopy, or cryo-EM. In cryo-EM, a protein is flash frozen onto a metal grid in a thin layer, ideally not much thicker than the diameter of the protein itself. Irradiating that layer with low-energy electrons rather than X-rays produces 2-D images of individual proteins. Thousands or even hundreds of thousands of these noisy images are then computationally sorted and reconstructed to build a 3-D image. Although X-rays yield cleaner images and can show how proteins bind small molecules, electron beams are less damaging to proteins. Cryo-EM also generates images from multiple copies of a protein, so an added benefit is that researchers can see how proteins might move or wobble in different conformations.
Advertisement
Over the years, computational power and microscope quality have gradually improved for cryo-EM, yielding higher- and higher-resolution biomolecule structures. In 2011, a breakthrough came when direct electron detectors became widely available. These devices provided big gains in the signal-to-noise ratio compared with previous indirect detectors, like charge-coupled devices. The advance triggered a flood of high-resolution cryo-EM structures determined by the scientific community. In addition to all these improvements, Butcher says, “the image processing methods have also taken leaps and bounds.”
Once derided as “blobology” for its blurry images, cryo-EM, whose pioneers won the Nobel Prize in Chemistry in 2017, is now churning out high-resolution structures of anything biologists can freeze to a grid. During the race to uncover the structure of SARS-CoV-2’s proteins, it’s been a standout.
We’re all in this together
That Rao’s and von Delft’s teams were able to share data about SARS-CoV-2’s protease to keep experiments running highlights another reason why structural biology research has advanced so rapidly during the COVID-19 pandemic. Even groups that were once competitors are working more closely with one another, according to scientists C&EN spoke with. One project that’s encouraging this type of collaboration is the COVID Moonshot crowdsourcing initiative, in which scientists across the world are being asked to help find inhibitors for the novel coronavirus’s main protease. The initiative took off after the contributions from Rao’s and von Delft’s teams. Von Delft’s group is a key contributor to the project.
And Rao and von Delft aren’t the only researchers sharing their work for the greater good. When C&EN spoke with McLellan in early April, he was still answering hundreds of emails a day from researchers worldwide, some wanting more details of his cryo-EM structure or asking him to share samples of the spike protein or the protein’s DNA code.
One partnership that struck up between McLellan and others recently bore fruit in the form of a publication. Collaborators led by Xavier Saelens at the VIB-UGent Center for Medical Biotechnology immunized llamas with SARS- and MERS-causing viruses to generate antibodies to the pathogens. Fusing a SARS antibody from a llama with a fragment of a human antibody yielded a hybrid that neutralized the virus responsible for COVID-19. The data suggest that such hybrid antibodies could be useful in combating coronavirus epidemics (Cell 2020, DOI: 10.1016/j.cell.2020.04.031). Llama antibodies are much smaller than human antibodies and are sometimes called nanobodies. Scientists have been trying to turn them into therapeutics for over 20 years, but it took until 2019 for the first nanobody-based therapeutic (caplacizumab) to be approved by the US Food and Drug Administration.
“Clearly, the whole world is looking for new entities at the moment,” says Ian A. Wilson at Scripps Research in California, who is using X-ray crystallography to look for antibodies that bind to SARS-CoV-2’s spike protein. Wilson’s focus since the 1980s has been using structural biology to develop universal vaccines for influenza and HIV.
The first labs to determine SARS-CoV-2’s protein structures may have had a background in coronaviruses, but other structural biology groups are now rolling up their sleeves to get involved too. To observe social-distancing guidelines while advancing science, many structural biology facilities are now open only to coronavirus-related projects and researchers.
“I’m not really surprised that so much activity is going on and so many structures are coming out,” Weiss says, pointing out that SARS-CoV-2 gives scientists a common foe to unite against. “I think the first instance when this happened was the race for the HIV protease in the ’80s and ’90s.”
With so many joining the fight, it’s hard to keep track of the structural research being done on SARS-CoV-2’s proteins, researchers say. New preprints—papers that haven’t yet undergone peer review—are coming out almost daily, most being posted on the server bioRxiv. And the scientists C&EN spoke with say they’re now relying on Twitter or their colleagues to alert them to new developments in the field.
“We all want it to move as quickly as we can,” Wilson says. “And I think things are moving at a very, very fast pace.” But developing any kind of drug or vaccine from these efforts and getting it through all the necessary steps required for regulatory approval will take time.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter