Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Microfluidic Device Purifies Nucleic Acids From A Handful Of Cells
Biological Analytics: Method could allow researchers and doctors to examine DNA and RNA from rare cells such as circulating tumor cells
by Erika Gebel Berg
September 25, 2013

When researchers study rare cells in the body such as those shed by tumors, they have precious little material to work with. Now, a team has developed a quick and easy microfluidic method for extracting DNA and RNA from small groups of cells—even a single cell—with minimal loss of material (Anal. Chem. 2013, DOI: 10.1021/ac402162r). The technique could help doctors diagnose specific types of cancer without doing invasive biopsies, the researchers say.
Circulating tumor cells are cells that a tumor sloughs off into the bloodstream. “We can collect these cells and perform the same genetic assays as you would with a biopsy,” without the need of an invasive procedure, says Lindsay Strotman, a graduate student in the laboratory of David J. Beebe at the University of Wisconsin, Madison. Analyzing the genetic material from the cells could help doctors select the appropriate course of treatment and reveal how mutations affect cancer cell behavior, she says.
To analyze these cancer cells, doctors and researchers want to identify mutations and measure expression levels for mutated genes. Both analyses require purifying and sequencing DNA and messenger RNA from the cells. Currently, a technician would do so using a commercial kit, which requires splitting the already tiny sample in two—one half for DNA extraction, and the other for RNA. When cells are scarce, that often leads to yields of DNA and RNA that are too small for further analysis. So Beebe’s team developed a microfluidic device that uses magnetic nanoparticles to capture DNA and mRNA from a single batch of cells.
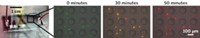
The microfluidic device has an input well, an oil well, and two output wells—one for DNA and one for mRNA, each on one face of the device. The input, oil, and output wells are connected by 0.3-mm-wide tunnels along either face of the device. The researchers add cells to the input well, along with a buffer that lyses the cells and magnetic nanoparticles coated with strands of nucleic acids that bind specifically to mRNA. After letting the mixture sit for five minutes, they drag a magnet across the device, pulling the nanoparticles through the tunnels from the input well, through the oil well, and into the mRNA output well. Cell debris doesn’t bind to the particles and can’t cross into the oil phase due to the oil’s hydrophobicity, so the debris is left behind in the input well. Only the mRNA-laden particles land in the output well. The researchers then add magnetic nanoparticles coated with silica to the input well. These beads bind DNA, so the researchers can then repeat the magnet pull to drop purified DNA into the other output well.
The researchers compared their device to a commercial kit, extracting DNA and mRNA from samples of one, 10, 100, and 1,000 prostate cancer cells. They measured the amounts of purified nucleic acid via quantitative polymerase chain reaction. The methods extracted similar amounts of mRNA for GAPDH, a highly expressed gene, but the microfluidic method purified more GAPDH DNA than the kit could. For example, the team’s device extracted 3.2 times more DNA from the 10-cell sample than the kit did. The kit took about an hour to do the extractions, and Strotman says their method takes about 10 minutes.
The researchers also demonstrated that their method produced usable material. They purified DNA and mRNA from circulating tumor cells provided by prostate cancer patients and then successfully amplified a cancer gene using the extracted nucleic acids.
The team took a laborious technique and made it easier for everyone, says Jeffrey J. Chalmers of Ohio State University. He’d like to try one of the devices in his own research on circulating tumor cells.




Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter