Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Fuzzy NMR Structures Get Sharper
Structural Biology: A computer program can refine NMR structures, making them as accurate as X-ray crystal structures.
by Erika Gebel Berg
January 27, 2014

For protein structure determination, X-ray crystallography typically beats nuclear magnetic resonance (NMR) spectroscopy in terms of accuracy. But a computer program called Rosetta may help NMR catch up, chemists report. They used the program to refine the NMR structures of a few dozen proteins and found that, in most cases, the resulting structures were as accurate as those determined by crystallography (J. Am. Chem. Soc. 2014, DOI: 10.1021/ja409845w).
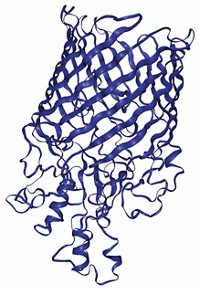
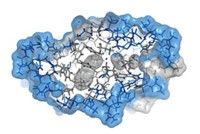
Knowing the precise structure of a protein is crucial to understanding its function. X-ray crystallography’s accuracy makes it the gold standard for solving protein structures. NMR is less accurate mainly because the technique analyzes proteins in solution where the protein’s atoms are freer to move around than in the crystalline state. This movement makes it difficult to pin down each atom’s position relative to the others in the protein.
Still, not all proteins are amenable to crystallization, leaving NMR as the best remaining option. Also, floating in solution could be a more realistic condition for proteins than sitting in a crystal lattice. “In some cases the NMR structure is more true to what’s in the cell,” says Gaetano T. Montelione of Rutgers University. For that reason, some chemists would like to improve the accuracy of NMR structures, he says.
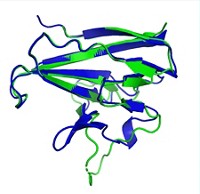
In the last several years, scientists have started using Rosetta to boost the quality of NMR structures. The program takes the relative positions of the protein’s atoms determined by NMR and then moves them slightly to refine the overall structure, basically transforming parts of the structure from a relatively high energy conformation to a relatively low energy one. Rosetta determines which conformations are high energy and which are low energy by applying basic physical chemistry principles and using information about stable protein folds found in the protein data bank, a digital database of known protein structures. Montelione and his colleagues set out to do the first general assessment of Rosetta refinement to see if it improved a range of NMR structures, using each protein’s corresponding crystal structure as a benchmark.
The researchers used 39 pairs of NMR and crystal structures of the same proteins as a test set. They looked at proteins between 5 and 22 kilodalton in size with varying topologies. The team applied the Rosetta refinement to each NMR structure and ensured that the tweaked structure still agreed with the NMR data collected in the lab. To assess whether the NMR structure got closer to the crystal structure, they performed a global distance test. This test compares the positions of corresponding carbon atoms in the two structures’ protein backbones and calculates whether the atoms are within a certain distance of each other. For 32 of the 39 protein pairs, the refined NMR structure was more similar to the crystal structure than to the unrefined NMR structure. Four of the remaining refined structures were equally similar to the unrefined and crystal structures. And three were slightly less similar to the crystal structure than the unrefined NMR structure was.
The results show the value of Rosetta, says Thomas Szyperski of the University of Buffalo. He thinks others in the field will pick up the method, possibly making it a standard for refining NMR structures.
Charles D. Schwieters of the National Institutes of Health questions whether the approach will apply to all NMR structures, because the ability of Rosetta to improve a structure may depend on how that structure was solved to begin with. There a lot of different methods to obtain NMR structures, he says.
Montelione next plans to test the approach on large proteins as well as other systems that tend to challenge structural biologists, such as membrane proteins and protein complexes.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter