Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Researchers Teach Old Microfluidic Chip New Tricks
Microfluidics: A commercial device for gene analysis becomes a platform for rapidly measuring binding affinity between proteins and peptide ligands
by Erika Gebel Berg
September 26, 2014

With a simple tweak, researchers turned a commercial microfluidic chip designed for nucleic acid analysis into a device that can rapidly measure binding affinities between proteins and peptide ligands (Anal. Chem. 2014, DOI: 10.1021/ac502605f). The approach can evaluate 2,304 binding pairs at a time using far less reagent than standard microarray-based methods.
William F. Burkholder of Singapore’s Agency for Science, Technology & Research (A*STAR) studies interactions between histones and regulatory proteins that control gene expression. The proteins can bind many different histone peptides, so Burkholder wanted the capacity to simultaneously screen multiple possible binding pairs. Existing methods have drawbacks, he says. Custom lab-on-a-chip devices need to be built from scratch, and microarray-based approaches require a lot of sample. Microfluidic chips cut down on sample volumes because they are closed systems, which eliminate evaporation, Burkholder says.
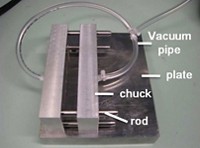
His team had started designing a new microfluidic chip for studying protein-ligand interactions when they realized they had a solution already in their lab: a chip made by Fluidigm. This platform, made for detecting specific genes through polymerase chain reaction (PCR), includes a microfluidic chip and dedicated fluorescence detector. One side of the chip contains 48 wells for DNA samples to analyze, and another side has 48 for primers of target genes. Microfluidic channels run from each well across the chip, creating a grid; a system of valves allows the samples and primers to mix and react at each of the 2,304 intersections without contaminating neighbors. The A*STAR team thought they could study 2,304 protein-peptide binding pairs by pipetting DNA regulation proteins into the sample wells and adding fluorescently labeled histone peptides to the primer wells.
During their proof-of-principle experiment, the researchers used fluorescence anisotropy to measure binding affinities. When a fluorescent peptide binds a protein, the rate at which the fluorescent peptide tumbles in solution falls, a change that is captured in the polarity of the emitted light. So the team placed a thin piece of polarizing film over the chip to filter the emitted fluorescent light on the basis of polarity on its way to the Fluidigm fluorescence detector.
The researchers wrote programs to calculate binding constants from the fluorescence anisotropy data they collected and found that their measured affinities squared well with values from the literature, as well as those they gathered using a standard 96-well microarray format. But the 96-well plate required 40 mg of protein, whereas the microfluidic assay needed only 40 μg to test the same number of interactions.
“The amount of reagent you save here is huge,” says John E. Scott of North Carolina Central University. This could make experiments feasible that would be too expensive or time-consuming if one had to buy or make protein for microarrays. Scott envisions applications in drug discovery, such as rapidly optimizing lead compounds by testing them against a target and its homologs.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter