Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Microneedle Patches Retrieve Multiple Biomarkers From Skin
Analytical Chemistry: Skin patches with arrays of microneedles could one day sample proteins for minimally invasive medical diagnostics
by Melissae Fellet
October 3, 2014

Patches containing arrays of microneedles can capture two specific proteins with high sensitivity from fluid in the skin of mice (Anal. Chem. 2014, DOI: 10.1021/ac5031682). The devices are first steps toward minimally invasive methods to detect multiple protein biomarkers without the need for drawing blood.
Doctors envision diagnostic tests to measure levels of particular biomarkers in blood to obtain information about a patient’s health or spot particular diseases early. Many diagnostic molecules, however, can be found in the interstitial fluid between skin cells. Researchers have built arrays of microneedles that painlessly pierce the skin to sample this fluid and detect small molecules and ions such as lactate, glucose, or electrolytes.
Protein biomarkers, however, are more difficult to sample this way because their concentrations in the skin are low, so researchers need to improve the sensitivity of their microneedle arrays to detect them. Directly capturing specific proteins with such devices would eliminate the purification steps typically needed for protein detection in blood samples.
In 2012, Simon R. Corrie of the University of Queensland, in Australia, and his colleagues developed a microneedle array that could retrieve a dengue virus protein from mouse skin (Anal. Chem., DOI: 10.1021/ac2034387). In the new study, they wanted to demonstrate that the device could retrieve more than one protein with high sensitivity and specificity, a crucial step toward making skin patches that can sample multiple biomarkers at the same time.
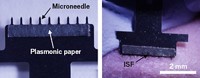
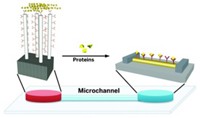
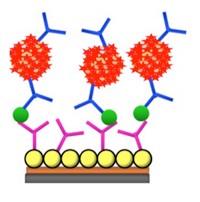
To make the device, the researchers etched a small silicon wafer to produce an array of thousands of 250-μm-long needles per square centimeter. Then they coated the needles with gold, followed by polyethylene glycol. Finally, the researchers attached antibodies that bind to HRP-2, a protein in the parasite that causes malaria. They optimized the conjugation conditions to maximize the number of antibodies on each needle.
The researchers also prepared two other arrays as controls, which could be important to calibrate detection in any skin patch used clinically. One array contained antibodies that bound immunoglobulin G (IgG), an immune protein abundant in skin. IgG captured by this array would confirm that the needles punctured the skin. The second array contained antibodies for the dengue virus protein, which should not be in healthy mice. Any protein attached to this array would be due to nonspecific binding.
The researchers applied the three arrays to the flanks of mice that had been injected with an engineered version of HRP-2. After removing the patches, they tested each array for the presence of protein, using an enzyme-linked immunosorbent assay.
The skin patches could detect the malaria biomarker when as little as 50 μg was injected into the mice’s tails. Moreover, the signals from the HRP-2 or IgG arrays were significantly higher than that from the dengue array, indicating that the HRP-2 and IgG arrays were sensitive and specific.
Ronen Polsky of Sandia National Laboratories thinks the study is impressive. Incorporating the positive and negative controls increases confidence in detection, and he thinks the next logical step would be to integrate the detection step into the device so that it does not have to be removed for analysis.
Corrie says the device is currently intended for single-use insertion and removal, though they also would like to incorporate detection methods into the device.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter