Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Peeking At A Cell’s Small Molecules
Mass spectrometry method can analyze an individual living cell’s metabolites and lipids in minutes
by Deirdre Lockwood
October 22, 2014

With a new mass spectrometry technique, researchers can survey the suite of small molecules inside a single cell by inserting a tiny probe. The method requires little sample preparation that would affect a cell’s chemistry and can analyze the contents of an individual cell in about three minutes (Anal. Chem. 2014, DOI: 10.1021/ac5029038).
Just as no two people are identical, no two cells are the same, says Zhibo Yang, a chemist at the University of Oklahoma who developed the new technique with his colleagues. Understanding biochemical differences between cells can help researchers pinpoint the hallmarks of cancer or neurological disease and investigate how individual cells respond to drugs. In 2012, the National Institutes of Health allotted $90 million to support research into this area through its Single Cell Analysis Program.
Historically, researchers have studied the appearance, modification, or disappearance of cellular biomolecules by breaking open large groups of cells into a lysate. This process yields an average signal that may not represent the conditions of small subpopulations or even individual cells. Yang and collaborator Anthony W. G. Burgett wanted to develop a way to screen individual living cells using mass spectrometry, which is one of the most powerful ways to study cellular small molecules, such as metabolites. Many other single-cell mass spectrometry techniques require putting cells under a vacuum or placing them into a matrix, which can kill or alter the cells.
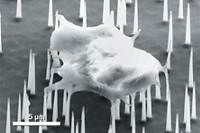
To develop the new technique, Yang made a tiny quartz needle with a tip less than 10 μm wide, smaller than most eukaryotic cells. Inside the needle are two channels; one provides a flow of solvent which pushes the contents of the cytoplasm through the other channel to a mass spectrometer for analysis. The scientists insert the needle tip into an individual cell with the help of a digital light microscope. Within seconds, the cell’s contents are sent to a nanoelectrospray ionization mass spectrometer for analysis. The full analysis of each cell usually takes three minutes.

One moment the cell is living in a culture, and in “the next instant its entire cellular contents are being sent to the mass spec,” Burgett says.
Using the method to analyze cervical cancer cells, the team identified several cellular metabolites and 18 different lipids that were known to exist in these cells. They also treated individual cells with anticancer drugs such as paclitaxel and detected the drugs and their metabolites in the cells.
“They’re extending mass spectrometry to a new dimension,” says Peter Nemes, a chemist at George Washington University who has developed a different approach for single-cell analysis. He says the new technique is impressive and will be easy to use.
Jonathan V. Sweedler, an analytical chemist at the University of Illinois, Urbana-Champaign, and editor-in-chief of Analytical Chemistry, points out that single-cell mass spectrometry could contribute to biological discoveries that are not feasible in single-cell analysis methods involving fluorescence imaging. Fluorescence techniques require knowing the molecular target beforehand. “When you only study preselected compounds, you may miss surprises,” he says. Burgett echoes this: Using the new technique, he says, “we’re not entirely sure what we might find.”




Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter