Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Structure Of The Measles Virus Nucleocapsid Reveals How It Wraps Up Its Genome
Structural Biology: Researchers tame the virus’s floppy capsid structure using cryo-electron microscopy
by Sarah Everts
April 16, 2015
| A version of this story appeared in
Volume 93, Issue 16

Cryo-electron microscopy has given scientists the first glimpse at how the measles virus wraps up its genome in its protective protein container. The 4.3-Å structure provides a platform for understanding viral replication and transcription and could enable the development of measles antiviral drugs, says Irina Gutsche, at the National Center for Scientific Research (CNRS), in Grenoble, France, who led the team of researchers.
The structure is also the first nearly atomic-resolution reconstruction of the full protein shell of an infamous family of negative-stranded RNA viruses that include rabies, mumps, and Ebola, comments David Bhella, a structural biologist at the University of Glasgow’s Centre for Virus Research.
The methodology used to solve this challenging structure is “an excellent starting point” for researchers trying to solve the overall structure of these other nefarious RNA viruses, adds Richard Plemper, at Georgia State University’s Institute for Biomedical Sciences, who was not involved in the work.
Like many viruses, the measles virus packages its genome into capsids formed by individual proteins that self-assemble into a large three-dimensional structure. But unlike most viruses, which have rigid capsids, measles and its relatives rely on a relatively flexible helical container. “Conceptually, the capsids are helical, but in reality they are more like floppy springs,” Bhella says.
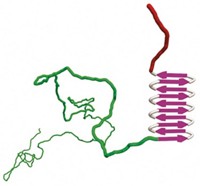
The flexibility comes from a disordered stretch of 150 amino acids in the capsid protein that serves as a docking site for the virus’s RNA polymerase. It’s this region that has made structural determination of these viruses so hard to tackle, Bhella says. Gutsche and her team chopped off that flexible region to wrestle the capsid into submission.
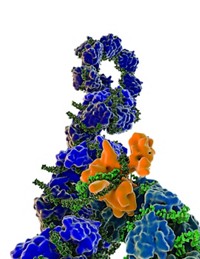
The structure reveals that the capsid has an interior helical core around which the RNA is wound (Science 2015, DOI: 10.1126/science.aaa5137). This core is formed primarily by the C-terminus domain of the capsid protein. The N-terminus domain of the capsid protein forms an exterior helical covering that protects the viral genome. Researchers wishing to design antiviral drugs may want to disrupt the assembly of this helical capsid, which seems to rely on the stacking of several aromatic side chains found in the N-terminus, Gutsche says.
With the structure in hand, Bhella says, many researchers will focus on how the virus’s RNA polymerase docks onto the helical capsid and how it pushes aside the capsid protein to gain access to the viral genome. Although the current structure provides a starting point for this work, he adds, it may require additional structural studies using the RNA polymerase and the full-length capsid protein.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter