Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Synthesis
Molecular Pump Mimics Natural Carrier Proteins
Molecular Machines: Synthetic system threads two rings onto a dumbbell-shaped rod
by Stu Borman
May 21, 2015
| A version of this story appeared in
Volume 93, Issue 21
Synthetic molecular machines that can shuttle molecules around could one day be used to create artificial muscles or transport molecules across membranes. Researchers have now brought such applications one step closer to reality by creating a molecular system that threads and traps large rings on a dumbbell-shaped molecular rod (Nat. Nanotechnol. 2015, DOI: 10.1038/nnano.2015.96).
J. Fraser Stoddart and coworkers at Northwestern University modeled this system on carrier proteins, which pump electrons, protons, ions, or molecules across membranes to produce concentration gradients in cells.
The work is “a fascinating example” of a nanoscale molecular system with a machinelike function, comments functional materials expert Ben L. Feringa of the University of Groningen, in the Netherlands. Although it is primitive compared with biological transporter proteins, it illustrates how an entirely artificial system can mimic key functions that have evolved in biological systems, Feringa says.
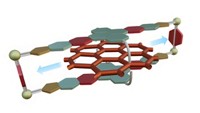
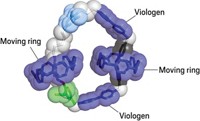
Stoddart’s team employed a clever design, says molecular machine specialist Jean-Pierre Sauvage of the University of Strasbourg, in France. It enables the pump to use redox reactions and heat to fish two cyclobis(paraquat-p-phenylene) rings out of solution and thread them, one at a time, onto a dumbbell-shaped molecule, where the rings become trapped in a high energy state. To start the pumping process, the ring in solution and viologen on the dumbbell are reduced, making them mutually attractive. The attraction pulls the ring over a dimethylpyridinium end cap on the dumbbell, and the ring forms a complex with viologen. Oxidation then returns the ring and viologen to their original states, making them repel one another and driving the ring toward an isopropylphenylene barrier. Mild heating enables the ring to slip over the barrier. This traps the ring, far from equilibrium, on an oligomethylene chain terminated with a diisopropylphenyl end cap. The second ring slips onto the dumbbell the same way. It takes about two hours to capture each ring.
Relative to previous work in the field, “the most remarkable improvement” of the new study is that it got two rings together onto a chain instead of just one, Sauvage says. Stoddart notes that the design is so versatile that threading more than two rings, and even an unlimited number, may eventually be possible.




Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter