Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Human Proteome Remapped
Antibody-based analysis allows protein mapping in individual tissues at the single-cell level
by Celia Henry Arnaud
January 26, 2015
| A version of this story appeared in
Volume 93, Issue 4
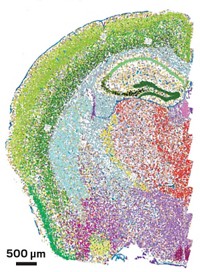
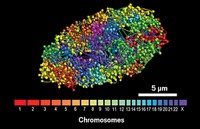
Independent teams announced in 2014 two draft maps of the human proteome determined using mass spectrometry methods. Swedish researchers have now acquired a more detailed map of the human proteome, this time using antibody-based analysis and RNA sequencing to achieve spatial mapping at the single-cell level (Science 2015, DOI: 10.1126/science.1260419). Mathias Uhlén of the Royal Institute of Technology and coworkers used 24,028 antibodies to map tissue-specific protein expression in 44 major tissues and organs. The team sequenced the transcribed RNA of 20,344 genes in 32 of the tissue types. Approximately 9,000 genes—the so-called housekeeping proteins—are expressed in all tissues, the team reports. In most tissues, only about 10% of the transcripts encode proteins that are elevated in that tissue relative to other tissues. Exceptions are the liver and the pancreas, in which 35% and 70% of transcripts, respectively, encode tissue-elevated proteins. The researchers analyzed subsets of the proteome, including secreted and membrane proteins. Only 677 putative protein-coding genes still have no experimental evidence, they note. These “missing genes” may turn out to not be protein-coding after all, the scientists say. Their data are available online in an interactive database at www.proteinatlas.org.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter