Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Method improves single-cell genome analysis
LIANTI leads to fewer errors that plague other amplification methods
by Celia Henry Arnaud
April 17, 2017
| A version of this story appeared in
Volume 95, Issue 16
Genomic changes in individual cells can eventually lead to cancer or other diseases. So scientists would like to be able to sequence the genome in a single cell. But the methods to do so can be plagued by the preferential amplification of some regions of the genome over others, leading to incomplete sequence coverage.
A team led by X. Sunney Xie of Harvard University and Peking University has developed a whole-genome amplification method that reduces such bias and errors (Science 2017, DOI: 10.1126/science.aak9787).
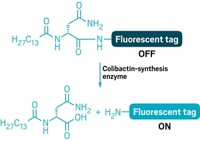
In the method, called LIANTI, researchers fragment genomic DNA from a single cell by inserting pieces of DNA called transposons. The transposons tag the DNA fragments so that they get amplified linearly instead of exponentially. The amplified DNA is then used to generate a library for subsequent DNA sequencing.
Compared with other whole-genome amplification methods, LIANTI has more uniform amplification and higher sequence coverage. The method enabled the detection of a type of mutation called copy-number variation, which involves the gain or loss of regions of the genome, which is hard to detect with high resolution using other amplification methods. The researchers were even able to characterize so-called micro-copy-number variations, which are smaller than 100,000 bases, with a resolution of about 10,000 bases.
Xie and coworkers used this ability to detect gains and losses of sequences to show that initiation of DNA replication is random and differs from cell to cell. They also showed that many single-nucleotide variations detected in previous single-cell sequencing are artifacts caused by instability of the DNA bases.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter