Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Thirty years of DNA forensics: How DNA has revolutionized criminal investigations
DNA profiling methods have become faster, more sensitive, and more user-friendly since the first murderer was caught with help from genetic evidence
by Celia Henry Arnaud
September 18, 2017
| A version of this story appeared in
Volume 95, Issue 37

The name Colin Pitchfork might not be as recognizable as Charles Manson or Jeffrey Dahmer. But for those in the forensic science community, it’s a name that holds weight. Pitchfork was the first murderer to be caught using DNA analysis.
When 15-year-old Dawn Ashworth was raped and murdered in Leicestershire, England, in late July 1986, Alec Jeffreys was a genetics professor at the nearby University of Leicester. A few years earlier, he had discovered that patterns in some regions of a person’s DNA could be used to distinguish one individual from another.
So far, Jeffreys had put his DNA pattern recognition technique to work in paternity and immigration cases, but now the police wanted him to help solve Ashworth’s murder as well as a similar one that happened in 1983. The police already had a suspect, Richard Buckland, who had even confessed to Ashworth’s murder. When Jeffreys analyzed DNA samples from the 1983 and 1986 crime scenes and from Buckland, he found matching DNA from both crime scenes—but the recovered DNA didn’t match Buckland’s genetic code.
In an attempt to find the real culprit—the one whose DNA had been left behind—the police undertook a genetic dragnet. They obtained blood and saliva samples from more than 4,000 men in the Leicestershire area between the ages of 17 and 34 and had Jeffreys analyze the DNA. They didn’t find a match until a man was overheard saying he’d been paid to pose as someone else and provide false samples. The person trying to evade the DNA dragnet was Colin Pitchfork. When Pitchfork’s DNA was analyzed, it matched the crime scene samples. Pitchfork was arrested on Sept. 19, 1987, and convicted and sentenced to life in prison the following January.
Although DNA evidence alone is not enough to secure a conviction today, DNA profiling has become the gold standard in forensic science since that first case 30 years ago. Despite being dogged by sample processing delays because of forensic lab backlogs, the technique has gotten progressively faster and more sensitive: Today, investigators can retrieve DNA profiles from skin cells left behind when a criminal merely touches a surface. This improved sensitivity combined with new data analysis approaches has made it possible for investigators to identify and distinguish multiple individuals from the DNA in a mixed sample. And it’s made possible efforts that are under way to develop user-friendly instruments that can run and analyze samples in less than two hours.
DNA profiling 101
A DNA profile is a list of numbers that indicate how many repeat units are in each copy of 20 marker regions located throughout the genome. Here’s how that profile is generated.
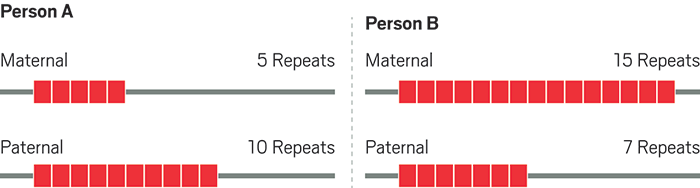
Chromosomes contain markers where short DNA sequences are repeated multiple times. The number of repeats at each marker varies from person to person, and each person has two copies, or alleles, of each marker, one inherited from their mother and one from their father.
To determine the number of repeats at each marker, forensic scientists extract DNA from cells in blood or other fluids or tissues, copy the DNA using the polymerase chain reaction, and separate the copied markers using capillary electrophoresis. The position of the peaks in the electropherogram correlates with the number of repeats in the two alleles for each marker. The electropherogram below shows the separation of five markers, including one where the number of repeats is the same in both alleles.

The resulting DNA profile for a person consists of the number of repeats in two alleles for each of 20 markers. Scientists enter DNA profiles into law enforcement databases as 20 pairs of numbers, such as 5,10 and 15,7.
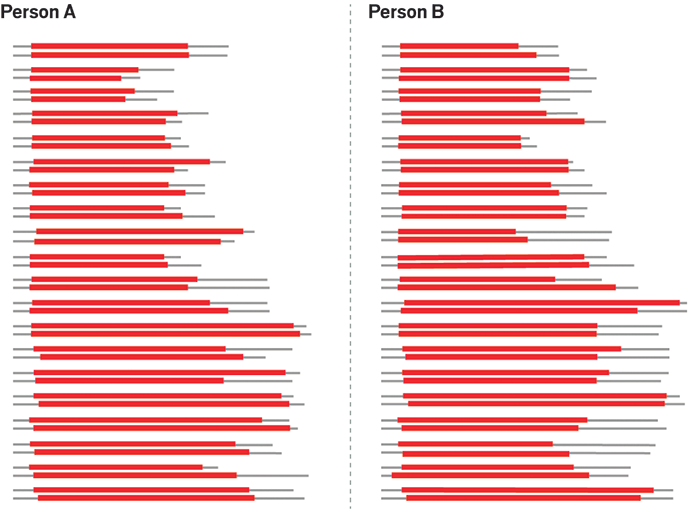
Then and now
DNA contains regions in which short sequences of bases are repeated multiple times. These repeats are found in many spots—or loci—throughout the genome. Because the exact number of repeats at any particular locus varies from person to person, forensic scientists can use these markers, called short tandem repeats (STRs), to identify individuals. What’s more, people inherit chromosomes from both of their parents, so individuals have two sets of STRs, each of which can have different numbers of repeats at the same locus. This pair of alleles can therefore provide even more specificity to a person’s DNA profile.
During DNA profiling, cells are collected and broken open to gain access to their DNA. Then forensic scientists copy the DNA regions of interest and measure the length of the repeat sequences at multiple loci. The length rather than the exact sequence of the repeats serves as a marker for DNA profiles because repeat length is sufficient for distinguishing among individuals. Although many STR loci dot the human genome, forensic scientists choose to analyze a small set of markers, rarely more than one locus per chromosome. Picking loci that are distant from one another ups the likelihood that the number of repeats at one locus is inherited independently of the number of repeats at another locus, thereby increasing the rarity of any particular DNA profile.
During the Leicestershire-area dragnet, Jeffreys used a type of repeat unit different from the ones used today. Those so-called minisatellites contained repeat segments that were dozens or even hundreds of bases long, says John M. Butler, special assistant to the director for forensic science at the U.S. National Institute of Standards & Technology. The overall DNA fragment at each locus could be tens of thousands of bases long, adds Tracey Dawson Cruz, a professor of forensic science at Virginia Commonwealth University (VCU).
Back then, forensic scientists like Jeffreys isolated and separated DNA fragments into size-dependent bands with gel electrophoresis. The DNA was labeled with radioactive phosphorus and detected with film that is sensitive to X-rays. “It took six to eight weeks to do a DNA analysis,” says Thomas F. Callaghan, senior biometric scientist at the U.S. Federal Bureau of Investigation Laboratory.
In the 1990s, the forensics community switched to STRs, which are a shorter type of repeat unit. The STRs used for forensics range from three to five bases long. Strung together with flanking sequences on either side, these STRs make up overall DNA fragments that are less than 500 bases long. The length of a DNA fragment correlates with the number of repeats it contains.
Rather than use X-ray-based gel electrophoresis, today’s forensic scientists measure the size of DNA fragments with a technique called capillary electrophoresis. Small fragments travel more quickly than large fragments through a gel-like material. As the separated DNA bits pass a fluorescence detector, they are registered as a series of peaks in an electropherogram. Capillary electrophoresis instruments are much faster, and they don’t require users to cast gels as is done in the X-ray-based gel electrophoresis technique of the past, VCU’s Dawson Cruz says. “The system is much more automated than before.”
While Jeffreys was developing his DNA fingerprinting method, Kary Mullis, who worked for the company Cetus, was developing the polymerase chain reaction (PCR), which would greatly advance DNA analysis techniques. In PCR, for which Mullis later won the Nobel Prize in Chemistry, enzymes “amplify” the amount of DNA in a sample by copying it many times, thereby making it easier to detect. Short pieces of DNA called primers identify specific regions of the genome and serve as starting points for copying them. The process involves repeated cycles of heating and cooling the sample. When carrying out DNA profiling today, forensic scientists use a different pair of PCR primers for each locus, so all the loci can be amplified in the same reaction without interfering with one another.
Before the boost in sensitivity provided by PCR, large samples such as bloodstains the size of a dime or a quarter were needed to get enough DNA for profiling, Butler says. With the advent of PCR and subsequent improvements, it’s now easier to get DNA profiles from much smaller samples. But what really made forensic DNA profiling take off was the creation of profile archives, Callaghan says.
The STR loci used in the U.S. Combined DNA Index System (CODIS) database are scattered throughout the human genome. The original 13 are highlighted in yellow, and the seven added in January 2017 are highlighted in green.

The rise of databases
“In the early days, a DNA case didn’t get worked unless there was a suspect and a blood sample from that suspect,” Callaghan says. In such cases, the payoff was obvious: The DNA could be used to include or exclude a suspect. Without a suspect, the benefit of analyzing crime scene samples wasn’t so obvious. They yielded little information without a suspect’s sample for comparison.
After governments started maintaining databases of DNA profiles, the incentive for running unknown samples skyrocketed. Violent crimes such as sexual assault and homicide have a high degree of repeat offenders, Callaghan says. So once databases were established, “if you ran a case where there was no suspect and you queried the database, you might be able to identify a suspect,” he says. Even if an unknown sample didn’t match an offender’s sample from the database, it might match DNA from other unsolved cases. In this way, serial rapists, for example, could be identified.
Database entries consist of a set of numbers that represents the summed-up STR repeats in each allele for a particular set of loci. In the U.S., DNA databases are part of the Combined DNA Index System, or CODIS, which was established in 1998 by the FBI. CODIS includes state databases and the national database known as the National DNA Index System (NDIS). Only accredited government laboratories—of which there are about 200—can submit profiles to NDIS.
For most of CODIS’s history, DNA profile entries have had to contain 13 loci. In January 2017, however, the number of loci for new CODIS profiles was increased to 20. The additional loci are primarily ones that forensic scientists in Europe were already using. Including those loci makes it easier to share DNA profiles internationally.
Any reference sample, which comes from a known suspect, added to the national database must have all the CODIS loci. Because such samples are fresh and come from a known source, “there’s no reason you can’t get 20 loci,” Callaghan says. “If you don’t get 20, run it again.”
There’s more leeway with crime scene samples, which are not as pristine. Labs must attempt to measure all 20 loci, but if they can’t get results for all 20, they can enter a partial profile into the database as long as it satisfies a statistical threshold prescribed by the CODIS search algorithm.
CODIS requirements dictate much of what forensic labs can do. The FBI approves commercial kits that contain the PCR primers and other reagents needed for amplifying the CODIS-designated loci.
To get that approval, manufacturers run developmental validation studies to make sure their kits work properly. Kits must be further validated by one or more approved CODIS laboratories to test the kits’ performance with regard to specificity and mixture accuracy. “We want to make sure that no matter which one of the 200 laboratories in the U.S. runs a sample, they get the same answer,” Callaghan says. Any lab that wants to use a kit must do its own internal validation.
Labs can use “home brew” kits if they want, but they must go through the same validation as commercial kits. “I’m unaware of anyone who’s not using a commercial kit,” Callaghan says. “The quality assurance and the validation and the expense and labor to establish something like that would be significant.”
Database requirements also constrain new technology. CODIS contains profiles of approximately 16 million convicted offenders and arrestees and 750,000 crime scenes. Any new technology must provide data that works with the existing database. “If there’s a faster, more sensitive way to do something, we would certainly be interested in that technology, as long as it’s backward compatible with what we’ve been doing with the national database since 1998,” Callaghan says.
The mixture challenge
The sensitivity of PCR allows investigators to generate DNA profiles from smaller samples than were possible in the early days of DNA profiling. The amount of DNA found in only a few cells is now enough to yield a profile, VCU’s Dawson Cruz says. But this improved sensitivity also has a downside. Today, analysis of a single sample is much more likely to lead to multiple DNA profiles because methods are sensitive enough to detect DNA that might have been in the background previously. For instance, a person may have touched a sampled doorknob before the criminal touched it. Teasing apart profiles from multiple contributors is complicated by the fact that PCR often produces so-called stutter peaks. For an allele with 10 repeat units, PCR amplification might drop or add a repeat, resulting in peaks that look like alleles with nine or 11 repeats. These stutter peaks are much smaller than the main peak. But stutter peaks from a major contributor—someone who left more cells behind—can be about the same size as main peaks from a minor contributor—someone who left fewer cells behind.
For most of the history of DNA profiling, analysts ignored this problem by using a threshold approach to determine which peaks from a capillary electropherogram to include in a profile. If a peak was larger than an experimentally defined value, it was included. If it was below that cutoff, it was left out because of the chance that the partner allele might be missing or because the peak might be confused with noise. In or out were the only options. That works fine with reference samples or single-contributor samples. But the analysis becomes much more complex if you want to identify a minor contributor in a mixture.
Advertisement
Today’s forensic scientists are moving away from this in-or-out approach. Instead, they are using mathematical methods that allow them to incorporate all the data in their analysis.
Software packages use algorithms to determine which combinations of DNA profiles better explain the observed data. “It turns out, of all the trillion trillion or so possible explanations, most of them don’t really explain the data very well,” says Mark Perlin, chief executive and chief scientific officer of Cybergenetics, the producer of TrueAllele, which was the first major statistical software for analyzing complex DNA evidence.
This mathematical approach to DNA data interpretation is known as probabilistic genotyping. The software proposes genotypes for possible contributors to a DNA mixture and adds them together to construct datalike patterns. The software gives higher probability to proposed patterns that better fit the data. A Markov chain Monte Carlo algorithm ensures a thorough search and finds explanatory genotypes.
For DNA analysis, “The power this change unleashes is truly staggering,” says John Buckleton, a representative of STRmix, the other major DNA mixture characterization software package. “It has turned the difficult samples into easy ones.” Analysts can now recover evidence from samples that they had previously declared inconclusive.
The new approach leads to better “match statistics.” Those statistics describe how much better a reference profile explains the evidence compared with a random profile. In the U.S., match statistics are a required part of any DNA-based testimony in court.
DNA evidence is no longer interpreted in ways to outright exclude individuals, says Bruce Weir, a professor of biostatistics at the University of Washington who focuses on DNA interpretation. “There could be a very low probability this person’s DNA is present in the sample, but it’s no longer zero,” he says. “That’s a profound switch in philosophy.”
Moving out of the lab
Not only has DNA analysis gotten faster since Jeffreys ran his first samples; it’s also breaking out of the lab.
The U.S. government’s Rapid DNA program aims to develop integrated systems that work without human intervention so that nontechnical personnel could use them in police stations to query databases. The program, which is being coordinated by the FBI’s Callaghan, is a joint project of the Departments of Justice, Defense, and Homeland Security. The Rapid DNA Act of 2017, which would allow DNA profiles generated outside accredited labs to be used to search CODIS, was passed by Congress and signed into law in August.
Under this law, in states with laws allowing arrestee testing, police can take cheek swabs at the time of booking. “You put a swab in. In 90 to 100 minutes, you generate a DNA profile,” Callaghan says.
The rapid DNA systems perform the same purification, amplification, separation, and detection steps that laboratories do. Software generates a profile that investigators can use to search the CODIS database for matches. Because the DNA from a reference sample comes from a single source, the system doesn’t need to perform mixture deconvolution.
Ande, formerly known as NetBio, is one of the companies that makes a rapid DNA system, also called Ande. Richard Selden, company founder and chief scientific officer, says the chemistry is the same as in conventional DNA analysis, but it’s much faster. The instrument decreased the time needed for PCR amplification from four hours to 17 minutes, he says.
Ande can be so much faster because it uses microfluidic chips and a very fast thermal cycler. With a typical PCR reaction, most of the time is spent ramping the temperature up and down. With the Ande system, “over the course of 15 minutes, you’re wasting a few minutes heating and cooling, but most of the time is the reaction being done,” Selden says.
The chip integrates all the steps of a typical DNA analysis. First, the cells are broken open and the DNA purified. Then the target loci are amplified. Finally, the amplified DNA is separated by electrophoresis and the sizes of the repeat segments determined. At the end of the process, about 90 minutes, the system automatically interprets the data to determine a profile, which is used to query CODIS or local DNA databases.
Advances on the horizon
As impressive as the current rapid DNA systems are, the forensics community is already thinking about the next generation of DNA analysis systems. Specifically, scientists are in the early stages of evaluating advanced DNA sequencing methods. In such methods, DNA sequences are analyzed by using arrays of single-stranded DNA fragments as templates for synthesis and detecting the order in which complementary bases are added. The next-gen methods have the advantage over conventional methods of being able to run many samples in parallel and thus being much faster.
Even though these new methods provide the DNA sequence, the size of the repeat regions can still be extracted from that sequence, so the methods should be compatible with existing databases.
That sequence can give forensic scientists other types of useful information when they’re searching for a culprit. “When you have hits in the database, it’s great because you have strong power of identification, and it’s well established in the courts,” says Susan Walsh, a forensic geneticist at Indiana University–Purdue University Indianapolis. “But what do you do when you have copious amounts of DNA at a crime scene but no comparative profile and no suspect?”
Walsh’s lab is working on ways to use DNA sequence information to establish key features of a perpetrator’s physical appearance. She is focusing initially on physical characteristics such as eye, hair, and skin color. “We find out through fundamental research where the genes related to physical appearance are,” she says. “Then we zoom in and design very small and sensitive assays to amplify those regions.”
Those assays identify single nucleotides at multiple locations throughout the genome. “To predict whether somebody has blue or brown eyes, we look at six pieces of DNA,” Walsh says. With kits designed for next-generation sequencing platforms, Walsh’s assays can be included in the same kits as STR profiling. “Not only do you get the STR profile, but you also get the most probable physical appearance profile,” she says.
Bruce McCord at Florida International University is studying whether epigenetic markers on DNA can be used as a forensic tool to determine a suspect’s age and the type of bodily fluid a sample came from. He and his colleagues use a modified PCR method followed by DNA sequencing to detect methylation differences in various kinds of tissue. “There are regions in your DNA where there are gradual changes in the level of methylation,” McCord explains. “That’s correlated with age.”
Although these methods hold promise, none of them has yet been approved for generating data to submit to CODIS, Callaghan says. “Next-generation sequencing holds the promise to be more discriminating and more sensitive than current capillary-based systems,” he says. The FBI is still evaluating whether next-gen sequencing provides results that are indeed compatible with existing databases, he says.
Forensic DNA analysis has come a long way since the Pitchfork case 30 years ago. There’s no telling where it will go in the next 30.





Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter