Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Polymers
Single-molecule experiment reveals polymer growth spurts
Researchers get the first detailed look at how catalysts crank out individual polymer chains
by Stephen K. Ritter
October 19, 2017
| A version of this story appeared in
Volume 95, Issue 42

When chemists think about polymerization, they typically envision a wormlike polymer growing smoothly and continuously from a catalyst. But the actual view of how polymer growth unfolds has remained murky because of the limitations of analytical techniques.
Using a pair of magnetic tweezers, optical microscopy, and spectroscopic techniques, Cornell University researchers led by Peng Chen, Geoffrey W. Coates, and Fernando A. Escobedo have achieved the first real-time visualization of single polymer chain growth. What they report is startling: Individual polymer chains don’t increase steadily but instead undergo consecutive wait and jump steps.
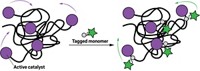
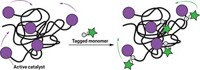
With the aid of molecular dynamics computer simulations, the researchers attribute this jerky mechanism to formation of polymer tangles—which they call hair balls—that form around the catalyst as thousands of new monomer units are added to the growing chain. The hair balls sporadically unravel after a couple of minutes, and a new hair ball starts to form.
Besides helping researchers better understand catalyst activity, polymerization rates, and bulk polymer properties, the researchers suggest their discovery of the growth spurts may be relevant to how cells produce biopolymers such as proteins, nucleic acids, and polysaccharides (Science 2017, DOI: 10.1126/science.aan6837).
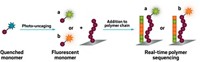
These new findings are “very cool,” says Suzanne A. Blum of the University of California, Irvine. Blum’s group has used fluorescently labeled molecules to study single-molecule dynamics and recently used this approach to watch catalytic activity as labeled monomers randomly got added to a growing polymer chain (Angew. Chem. Int. Ed. 2017, DOI: 10.1002/anie.201708284).
The newfound wait and jump steps of polymer lengthening were previously obscured by “ensemble averaging,” Blum explains, in which researchers use techniques such as dynamic light scattering to observe all the molecules in a sample at once, and information about size distributions and other polymer parameters are extracted from the data. Single-molecule approaches avoid limitations of ensemble averaging, Blum says, but these measurement techniques are often a double-edged sword because the resulting data are a challenge to interpret without corroborating spectroscopic methods. By including molecular dynamics simulations, the Cornell team was able to get a clearer picture of conformational changes in the growing polymer.
“The ability to see dynamics in an important reaction like polymerization and to understand them through modeling is an exciting technological advance,” Blum says.
In their single-molecule experiment, the Cornell researchers attached the free end of a polymer chain to a glass surface using a silane linkage and attached the ruthenium catalyst at the growing end of the polymer to a magnetic particle held in place by a pair of magnetic tweezers. By tracking the position of the magnetic particle, the team achieved real-time visualization of a single chain’s growth during ring-opening polymerization.
“This is just a superb piece of science,” says Craig J. Hawker of the University of California, Santa Barbara. Hawker’s group recently reported the one-pot synthesis of block copolymers using five different monomers with widely varying properties (Angew. Chem. Int. Ed. 2017, DOI: 10.1002/anie.201707646). The new polymer-growth monitoring process could help researchers understand how to better control such exotic polymerizations and allow them to tune the macroscopic properties of polymer networks to design new functional materials, he says.
“The Cornell team’s tantalizing view of how polymer chains grow sheds light on many synthetic challenges dating from the earliest days of polymer chemistry,” Hawker adds. “This work will have major implications far beyond single-chain dynamics.”
This article has been translated into Spanish by Divulgame.org and can be found here.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter