Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Materials
Engineered myosin motor uses RNA arm to march on protein fibers
Researchers can control the direction of the motor’s movement with strands of DNA
by Stu Borman
November 9, 2017
| A version of this story appeared in
Volume 95, Issue 45

Muscles contract thanks to the work of a protein called myosin. Using energy from adenosine triphosphate, this molecular motor works with fibrils of the protein actin to drive muscle contraction. Myosin can also move along an actin filament by attaching to actin, swinging a pivoting arm to pull on the filament, detaching from actin, and repeating the process.

Previously, researchers have re-engineered myosins into motors that change direction on actin filaments in response to ions or flashes of light. And others have created artificial biomolecular motors made completely from DNA.
A research team including associate professor of bioengineering Zev Bryant and postdoc Tosan Omabegho of Stanford University has now created a hybrid motor in which RNA components replace myosin’s pivoting protein arm (Nat. Nanotechnol. 2017, DOI: 10.1038/s41565-017-0005-y). The protein-RNA hybrid moves faster than previous re-engineered protein motors and artificial DNA motors, and the researchers can change its direction of motion by adding strands of DNA with specific sequences, potentially a more versatile form of signaling than the use of ions and light.
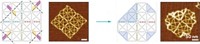
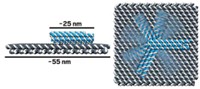
The team tested two versions of the motor. They fixed monomeric forms of the motor to surfaces and watched the hybrids move actin filaments. They also developed a tetrameric form that can walk along actin filaments that are fixed to a surface. They formed the tetramer by generating a cyclic DNA complex that binds to the motors’ RNA arms, enabling four motors to arrange themselves symmetrically around the ring.
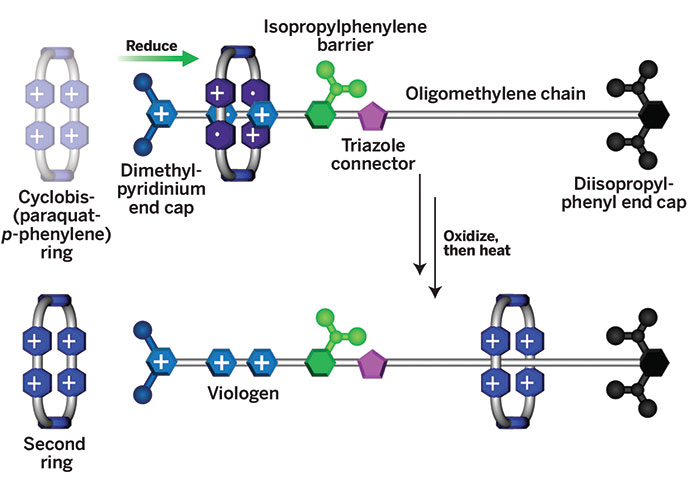
To change the direction the motor moves, the researchers add a DNA strand called a switch strand, which binds to a switch loop, an RNA domain at the base of the motor’s arm. Addition of the switch strand causes the RNA arm to undergo a conformational change that reverses the direction of the arm’s movement and, therefore, the direction of the motor’s movement. If the scientists then add another piece of DNA called a switchback strand, it base-pairs with the switch strand and removes it from the arm, returning the motor to its original direction of motion. Switch loops and DNA strands customized with different sequences make it possible to control distinct motors to perform different tasks. Such versatility of control is much harder to achieve with ion- or light-based directional switching methods, the researchers say.
“To my knowledge, this is the first demonstration of using nucleic acids to control the directionality of a single-protein molecular machine,” comments biomolecular mechanics expert Rizal Hariadi of Arizona State University. “It is the most impressive molecular engineering of controllable bidirectional molecular machines I have seen.”
In the system’s tetramer form, the motors work together to walk along a fixed actin filament at speeds of 10 to 20 nm per second. This is slower than many native myosins, which have benefited from billions of years of evolution and can move at up to micron-per-second speeds. But the tetramer’s speed is about 10 times as fast as earlier engineered protein-only bidirectional motors and about 100 times as fast as previous artificial DNA-only motors.
The researchers think a potential application of the motors is to serve as molecular transporters. Like the brooms fetching pails of water in “The Sorcerer’s Apprentice,” they might drop off molecular cargo at a defined location and then go back to pick up more cargo. The team also envisions producing versions of the hybrid motors that could be encoded genetically so living cells could synthesize them to perform intracellular tasks.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter