Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Chemists engineer bacteria to use unnatural DNA bases to make unnatural proteins
Living microorganisms transcribe and translate DNA with six bases instead of the usual four
by Stu Borman
November 30, 2017
| A version of this story appeared in
Volume 95, Issue 48

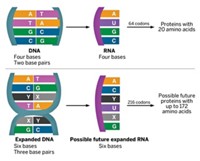
The genome of every cell on Earth uses four DNA bases—adenine, thymine, cytosine, and guanine—to encode proteins. Chemists have long dreamed of expanding that set to create cells that work with both natural and unnatural nucleic acids. An expanded collection of bases could allow cells to synthesize unnatural amino acid-containing proteins that have new functions and could be useful as novel drugs, vaccines, and nanomaterials. But the closest researchers had gotten was to translate DNA with expanded base sets into proteins in test tubes.
Now, Floyd E. Romesberg of Scripps Research Institute California and coworkers have gone all the way by putting an expanded genetic code into bacteria and getting the microbes to use it to synthesize an unnatural protein (Nature 2017, DOI: 10.1038/nature24659).
The researchers got a strain of Escherichia coli to work with dNaM and dTPT3, aromatic DNA bases that pair with each other through complementary packing and hydrophobic forces instead of the usual hydrogen bonding that natural bases use. They engineered the bacteria with a circular DNA plasmid that had a gene for green fluorescent protein (GFP). Inside that gene, the researchers inserted the unnatural base pair dNaM-dTPT3 at a site where they wanted to incorporate an unnatural amino acid in GFP.

The scientists modified the bacteria to express an enzyme that transcribes the GFP gene to a messenger RNA with the RNA version of NaM at the corresponding site. To allow the bacteria to translate mRNA containing this unnatural base, the researchers also engineered the cells to produce a transfer RNA that could recognize NaM. During protein translation, in general, a three-base anticodon in a tRNA binds to a three-base codon in an mRNA, causing the ribosome to add an amino acid that is bound to the tRNA to the growing protein chain. In the new study, the engineered tRNA contained the RNA version of TPT3 in its anticodon to pair with NaM in the mRNA.
The scientists tested the system with two different tRNAs, each carrying a different unnatural amino acid—an N-propargyl-lysine or p-azido-phenylalanine—and demonstrated that the bacteria added each at the proper spot in GFP. Romesberg earlier cofounded the biotech firm Synthorx to commercialize this type of expanded-DNA technology.
The work shows that unnatural base pairs are compatible with the molecular biological machinery inside cells, “opening the door to a new stage of genetic alphabet expansion,” comments Ichiro Hirao of A-STAR Institute of Bioengineering & Nanotechnology.
Steven A. Benner of the Foundation for Applied Molecular Evolution is skeptical of how broadly applicable this technology is. He believes hydrophobic base pairing may work only when unnatural bases are sandwiched between natural ones for stabilization, and he doubts this type of base pairing is a credible system for sustaining varied forms of semisynthetic life.
On the other hand, Dieter Söll of Yale University thinks the work has great promise. “It is the first real proof that living things will be able to make customized proteins with multiple unnatural amino acids in the future,” he says.
This article has been translated into Spanish by Divulgame.org and can be found here.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter