Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Gene Editing
Light-controlled guide RNA hones CRISPR targeting
Technique modifies subsets of cells and frees gene editing proteins for potential further improvements
by Jyoti Madhusoodanan, special to C&EN
May 5, 2020

Controlled gene editing holds the potential to transform disease treatment, crops’ climate tolerance, and our ability to investigate genetics. The CRISPR-Cas9 gene editing system has proved a key advance because of the ease and precision of making targeted genetic changes. Now, two independent teams demonstrate that CRISPR-Cas9 can be turned on in only a subset of cells in culture and in zebrafish embryos at specific times, by controlling the system with light more tightly than previous attempts (ACS Cent. Sci. 2020, DOI: 10.1021/acscentsci.9b01093; Angew. Chem. Int. Ed. 2020, DOI: 10.1002/anie.201914575). Unlike earlier efforts, the teams’ method targets a part of the CRISPR-Cas9 system known as the guide RNA.
Light offers many advantages as a switch for controlling gene editing in cell culture or tissues: it is minimally invasive, works quickly, and unlike biochemical controls, doesn’t need additional chemical agents added to cells. The ability to manipulate where and when genes are edited could help researchers target subsets of cells within an organ, or only manipulate a gene at certain times during embryonic development. Researchers have already developed a variety of light-activated CRISPR-Cas9 systems, all of which involve light-controlled Cas9 proteins. But it’s hard to make additional upgrades that would improve gene editing selectivity or efficiency to light-controlled proteins. So it would be better to add the light control elsewhere to allow researchers to more freely engineer Cas9, says Alexander Deiters of the University of Pittsburgh, who led one of the studies.
So the teams focused on a part of the system called the guide RNA, which is designed to bind to the DNA sequence that is targeted for cutting, creating a DNA-RNA complex that the Cas9 enzyme recognizes and snips. In the new studies, teams led by Deiters and Molly M. Stevens of the Karolinska Institute modified guide RNA molecules so they carried light-sensitive molecules that “caged” the RNA to prevent DNA pairing. When exposed to the right wavelength of light, the caging molecules were cleaved off and the RNA could bind to the target DNA, initiating the gene editing process.
“They essentially install this nice photo-cleavable knob on the RNA bases involved in the Watson-Crick pairing” that’s necessary for Cas9 to cleave DNA, said chemical biologist Amit Choudhary of the Broad Institute, who was not involved with either study, speaking of the Stevens group’s work. “In the absence of light, the pairing doesn’t happen and there’s very low background activity.”
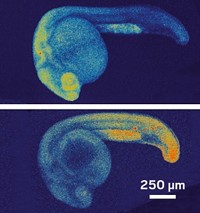
The researchers treated cultured human cells with the photocaged RNAs, and found no signs of DNA editing in the absence of light. When illuminated, the activated RNA restored 70% of gene editing activity seen with control RNA molecules that lacked the photo-active attachments in the Stevens group’s experiments. Both groups tested their systems in zebrafish embryos and found they could introduce mutations in one eye and not the other by illuminating it. Similar results from two independent studies such as this are “particularly reassuring” that the method will prove reproducible, Lapatrada Taemaitree and Tom Brown of the University of Oxford wrote in a perspective (ACS Cent. Sci. 2020, DOI: 10.1021/acscentsci.0c00350). “In principle, multicolor targeting systems could be developed to allow different genes to be switched off simultaneously or at different times.”


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter