Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Microbiome
Researchers elucidate colibactin structure
The team synthesized the molecule, confirming its structure and providing a route for studying the difficult-to-isolate bacterial metabolite
by Celia Henry Arnaud
August 9, 2019
| A version of this story appeared in
Volume 97, Issue 32

Some Escherichia coli bacteria in the gut produce a family of metabolites called colibactins, which are associated with increased risk of colon cancer in people. But for more than a decade after its discovery, researchers couldn’t isolate colibactin and had no clue what it looked like.
Two teams have previously reported structures of DNA-colibactin adducts (Biochemistry 2018, DOI: 10.1021/acs.biochem.8b01023; Science 2019, DOI: 10.1126/science.aar7785), but they didn’t figure out the whole structure of colibactin. Now, one of those teams—Jason M. Crawford, Seth B. Herzon, and coworkers at Yale University—has deduced the structure of the major colibactin metabolite from a cross-linked adduct of the molecule with DNA (Science 2019, DOI: 10.1126/science.aax2685). In the process of determining the structure, the researchers developed a synthesis that will make future studies of the molecule easier.
“With these latest results from two independent labs, colibactin structure is now resolved,” says Jean-Philippe Nougayrède, who studies pathogenic and commensal bacteria at Inserm, a French medical research institute. “Its structure and mode of action fit perfectly with its genotoxicity that has been documented in vitro, in cultured cells, and in vivo, in mice and rats.”
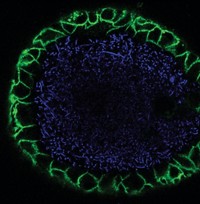
To elucidate the major colibactin structure, the Yale team captured the molecule as a cross-linked adduct with two adenines in DNA. It then used a combination of mass spectrometry, isotope labeling, and chemical synthesis to determine the structure.
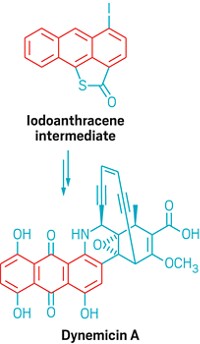
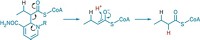
Colibactin includes two electrophilic spirocyclopropyldihydro-2-pyrrolones and an easily hydrolyzed dicarbonyl. It attaches to DNA through the opening of its cyclopropane rings. The team synthesized colibactin via acylation of a diamine with two β-ketothioesters, followed by double cyclodehydration.
“The latest results put an end to the colibactin mystery,” Nougayrède says. “We can now focus on other important questions—its role in bacterial virulence, commensalism, and cancer.”


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter