Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Antibiotics
Bacteria make a meal of penicillin
Engineered bacteria could be used for bioremediation of antibiotic-contaminated soils
by Celia Henry Arnaud
May 7, 2018
| A version of this story appeared in
Volume 96, Issue 19

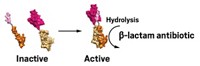
Somewhere in the soil microbiome, bacteria are chowing down on penicillin and other β-lactam antibiotics. A team of researchers led by Gautam Dantas of Washington University in St. Louis provides evidence for a pathway those bacteria might use to incorporate β-lactams into the microbes’ central metabolism (Nat. Chem. Biol. 2018, DOI: 10.1038/s41589-018-0052-1). After genomic and transcriptomic analysis of bacterial strains that can survive using penicillin as their sole carbon source, the researchers propose that the bacteria first cleave the β-lactam ring to produce benzylpenicilloic acid. That compound acts as a substrate for an amidase enzyme, which hydrolyzes the amide bond to release the phenylacetic acid side chain. Phenylacetic acid can be metabolized into acetyl-CoA and succinyl-CoA, which feed into the bacteria’s central metabolism. The researchers engineered a set of these penicillin-processing enzymes into Escherichia coli, creating a strain that subsists on penicillin as its sole carbon source. They propose that such engineered bacteria might be used for bioremediation of antibiotic-contaminated soils. But the researchers acknowledge that such bioremediation comes with the risk of spreading the resistance and degradation genes to other organisms.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter