Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Drug Development
Drug hunters explore allostery’s advantages
After decades of focus on proteins’ active sites, a new crop of biotech firms is shifting attention to proteins’ hidden regulatory pockets
by Lisa M. Jarvis
March 10, 2019
| A version of this story appeared in
Volume 97, Issue 10

Credit: Chris Gash; Hotspot Therapeutics (protein)
In brief
After decades focused on the active site of proteins, drug hunters are increasingly turning their attention to allosteric sites—the often tough-to-find regulatory pockets. A crop of allostery-focused biotechs has emerged to illuminate proteins’ hidden sites with a combination of better experimental and computational tools. After years of finding allosteric modulators by accident, chemists think they have the tools to rationally design them. Read on to understand why this approach might crack open difficult drug targets and lead to safer, more effective medicines.
Drug hunters have spent years cutting chemical keys to fit the most obvious biological locks: a protein’s active site, the spot where its natural substrate clicks in to turn it on or off. Walk into any big pharma company lab, and you will find the first step of that process: crane-like robotic arms pipetting tiny drops of compounds onto well plates full of target proteins that are then zipped across the room for analysis. These ubiquitous high-throughput screens are all in the service of finding hits, which more often than not are defined as molecules that bind a protein’s active site.
But there are other locks to pick. Allosteric sites—grooves beyond a protein’s active, or orthosteric, site—can be tough to find, and matching them with the proper key is a more nuanced exercise. Yet if discovered, they can allow chemists to develop safer medicines and tackle long-elusive targets. The allure is prompting more biotech and big pharma firms to flex their lock-picking skills.
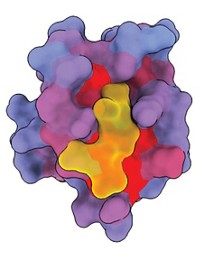
The concept of allostery isn’t new, but most allosteric drugs in the clinic or on the market were found by accident rather than by design. That’s because scientists were long stuck with a simple snapshot of their target, often just a close-up of the active site, when what they needed was to see the protein in its full, dynamic, shape-shifting glory. When a partner approaches, a protein might raise an arm or twist and bend, a conformational change that opens up hidden pockets.
Drug-discovery scientists are getting better at seeing and deciphering the function of these hidden sites.
The problem in the past was a lack of tools and drive on the part of researchers, says Nick Keen, chief scientific officer at Bicycle Therapeutics, who previously worked on allostery at Novartis. “I think people honestly followed the path of least resistance. And that’s really changed.”
Researchers now have new technologies for finding proteins’ hidden pockets, cheaper ways to measure the biophysical properties of proteins, and access to previously unfathomable computational power to model how proteins change in the presence of a drug.
Those advances coincide with a clear business rationale for companies to move out of their active-site comfort zone. “The tree’s been sort of picked,” says Mark Goldsmith, CEO of Revolution Medicines. Most of the easy-to-drug proteins have been or will soon be tackled, pushing companies to take on targets that previously proved elusive.
With the scientific and business cases converging, a wave of young biotech firms has emerged to take on the challenge of allostery. Their scientific approaches vary, as do the diseases they want to conquer, but they’ve all amassed significant funding to try to unlock those hidden pockets. Now, they just need to show that their technology can live up to the hype.
Keys to the kingdom
Beyond their need for new ways to access tough drug targets, researchers have many other reasons to embrace allosteric modulation. The most obvious is selectivity: a given protein’s active site can look awfully similar to that of its family members, making it difficult to design a compound that doesn’t unlock them all. An allosteric site is more specific to an individual protein, potentially leading to safer medicines.
Similarly, the active sites of healthy and mutated proteins usually look alike, meaning conventional drugs shut down the good along with the bad. Allosteric modulators offer the potential to spare the healthy proteins.
“The allosteric site has a promise of selectivity that’s very difficult to ignore,” says Larry Burgess, head of chemistry at Vividion Therapeutics. With the human proteome spanning some 20,000 proteins, companies long ago gave up on being able to fully predict the off-target effects of their active-site compounds, he says. “If you can fill a site that is highly unique, the idea is you’ll be highly selective.”
Moreover, jamming a compound in a protein’s active site turns it off. But what if instead of inhibiting it, you want to just dampen its effect or even turn it on? For that matter, what if the active site has been mutated or was never nice and welcoming to begin with?
In theory, allosteric sites can solve all those drug-discovery quandaries. They introduce the possibility of fine-tuning a protein’s activity—one modulator might shift a protein into a conformation that shuts it down, while another might amp up its activity—and accessing drug targets that are inaccessible to active-site inhibitors.
“You can view an allosteric binder as a knob where you can dial in how much activation or inhibition you want,” says Dorothee Kern, a biochemist at Brandeis University, a Howard Hughes Medical Institute investigator, and cofounder of Relay Therapeutics. That rheostat “is already there. Nature has built these allosteric networks in for regulation. Let’s take advantage of them.”
Indeed, allostery’s assets are thanks to nature’s need to carefully manage the bustling activity inside every cell. “Nature really only has a handful of chemistries that are performed—oxidation, reduction, phosphorylation, ubiquitination, etc. You can count them on two hands,” says Gerry Harriman, cofounder and CSO of allostery-focused HotSpot Therapeutics. “Yet it has thousands of proteins in a cell and has to control them exquisitely—otherwise there’s cellular chaos.”
That complex cellular orchestration is enabled by regulatory hot spots—allosteric sites that nature has tucked into proteins in order to control their activation. “Function and regulation using these hot spots has to be controlled very specifically,” Harriman says, and that means the sites tend to be different from one another and from active sites.
Proteins’ secret surfaces
All those advantages suggest that an allosteric site is a pretty rational starting point for drug hunters. “If you’re starting with a clean slate and you’re looking at affecting the activity of any given enzyme, allosteric inhibition is probably the way to go,” agrees Bicycle’s Keen, who has served as a consultant for HotSpot.
And yet the industry spent decades focused instead on protein active sites. The main reason was that active-site kinase targets were numerous and easy to drug. Big pharma firms were also under the spell of their massive compound collections and high-throughput screening strategies, which were tailored for finding active-site inhibitors.

In addition, companies were burned by failures in the allostery space. Craig Lindsley, codirector of the Vanderbilt Center for Neuroscience Drug Discovery, says industry didn’t devote the time and resources needed to truly unravel the biology of a protein. They wanted to “jump in, get to an assay, and go,” says Lindsley, who has spent his career working on allosteric modulators in both industry and academia. “Allosteric modulation requires more up-front investment.”
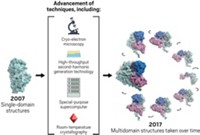
One stumbling block was that scientists didn’t have the best tools to explore proteins’ hidden topographies. That situation has changed in the past five years with advances in structural biology and cheaper tools for “hard-core biophysical measurements,” Keen says. X-ray crystallography, cryogenic electron microscopy, room-temperature crystallography, and screening technologies that capture conformational changes in a protein have all been pivotal in seeing and understanding allosteric pockets.
Those protein images are enhanced by shuffling between experimentation and computation, which itself has undergone a boom. “You can actually do drug design right on your cell phone today,” Revolution’s Goldsmith marvels.
Researchers have the processing capability to improve algorithms for describing the interactions between proteins and small molecules. “We now have a much better understanding of the dynamics of proteins,” says Ramy Farid, CEO of Schrödinger, a computational-chemistry company whose platform is used by many biotech firms working on allosteric modulation. And computational firms have improved their understanding of how water behaves around proteins, which is critical for calculating the strength of interactions between proteins and their ligands.
Farid cautions that computers are not yet sophisticated enough to reliably identify allosteric pockets. But many researchers are nonetheless encouraged by the potential of the parallel revolutions in molecular modeling and experimental tools.
In fact, they are already revealing allosteric sites on long-sought-after drug targets. In the past three years alone, multiple allosteric sites on G protein–coupled receptors have been published, says Jon Mason, a senior research fellow at Sosei Heptares who is also a biotech consultant. “This is my 40th year in the pharma industry, and now we can finally do things rationally.”
And success stories are emerging. One fruitful area is phosphatases, enzymes that regulate cell signaling by plucking phosphate groups from proteins. Companies spent years trying to develop active-site inhibitors for phosphatases, but they proved elusive.
One problem was the similarity between family members’ active sites, which made it “particularly difficult to engineer any degree of selectivity,” Revolution’s Goldsmith says. And the enzymes’ shallow, highly polar active sites are inhospitable to the kind of small molecules that can be turned into pills, causing many phosphatase-inhibitor programs to crash and burn.
But a combination of new technologies, better images of certain targets, and a shift away from high-throughput screening to whole-cell screening led to a small crop of allosteric phosphatase inhibitors.
One of the most prominent phosphatase projects came out of Novartis, which in 2016 revealed the structure of the first allosteric SHP2 inhibitor. Mutations in the gene that codes for SHP2 have been linked to a variety of cancers, while SHP2 also enables players in the cancer-associated Ras-MAP signaling pathway.
Novartis scientists used clever screening approaches to come up with an allosteric SHP2 inhibitor that is now in a Phase I study. The company subsequently disclosed an allosteric BCR-ABL inhibitor that, along with the SHP2 inhibitor, “sort of unleashed the floodgates,” Keen says.
Indeed, with Novartis’s crystal structure of the SHP2 allosteric site also in the public domain, work on the target flourished. Revolution recently began a Phase I study of RMC-4630, an allosteric SHP2 inhibitor that last year it licensed to Sanofi for $50 million. Revolution’s portfolio also includes an earlier-stage SHP1 inhibitor.

Locksmiths abound
Phosphatases aren’t the only tough targets being toppled. Nimbus Therapeutics developed an allosteric inhibitor of acetyl-CoA carboxylase (ACC), an enzyme that had long eluded researchers. Medicinal chemists just couldn’t come up with a small molecule to potently block ACC’s shallow, hydrophobic active site.
Nimbus looked to nature for inspiration. It turned out that a bacteria-made macrolide called soraphen A combats yeast by clinging to an allosteric site on ACC. The biotech and Schrödinger, its computational partner, probed the interaction to generate virtual ACC inhibitors that Nimbus’s chemists eventually turned into a drug candidate.
The discovery process took just 18 months, and the drug candidate, firsocostat, was so promising that, in 2016, Gilead Sciences paid $400 million to license it from Nimbus. The compound is currently in Phase II trials to treat people with a serious liver disease called nonalcoholic steatohepatitis.

Such successes have convinced investors that allostery is worth revisiting. In less than a year, more than half a billion dollars have flowed from venture capital and big pharma companies into young biotech firms working on the concept.
Each company has its own angle on allostery. Nimbus’s ACC program led a handful of the firm’s executives to consider broadening the concept of using nature as a guide to a protein’s regulatory pockets. Last year, they formally launched HotSpot with $45 million in venture capital money to do just that.
One of HotSpot’s first tasks was to explore, computationally, the surfaces of thousands of proteins to identify hidden pockets that researchers then try to characterize.
“We’ve come up with 50 or so different descriptors that relate to regulatory pockets,” CEO Jonathan Montagu says. “It’s a bit like when chemists look at active sites: they know exactly what they’re looking for because they’ve been studied extensively. We’re coming up with those same rules now for regulatory pockets.”
Those descriptors are fed into a machine-learning model that scores unknown pockets according to their likelihood of being a regulatory hot spot.
While HotSpot focuses on finding and exploiting regulatory sites, Relay describes its approach to allosteric modulation in more biophysical terms. “In textbooks, the drug simply binds the protein, and then, game over,” Brandeis’s Kern says. But in reality, binding is a series of conformational changes, with the compound changing the energy landscape of a protein. The idea is that different compounds can cause different changes. One molecule might shift the protein in a direction that shuts down the active site, while another might turn it on, Kern explains.
Relay is combining experimental biophysical techniques—for example, specialized screening methods that allow researchers to detect conformational changes that occur when a molecule interacts with a target—with molecular dynamics simulations to understand a protein’s movements. The goal is to develop small molecules that, by binding an allosteric site, can push a bad-behaving protein back into a healthy state. Although Relay has yet to reveal the cancer drug targets it is chasing, the firm has raised $520 million to develop its protein-motion-based platform.
Vividion is taking yet another approach to finding hidden pockets on proteins. Its researchers wash chemical probes over live cells and then look for probes that have bound to cysteines on the surfaces of proteins. The method lets the company map all the nooks and crannies on a protein’s surface.
Because the experiment is meant to find only molecules that bind to proteins, not ones that actually change protein function, the researchers “are able to see pockets in proteins that have never been found before,” says Vividion CEO Diego Miralles. The next step is determining whether that interaction produces a change in biology.
Not all the drugs that are developed from those screens will be allosteric modulators. Still, Miralles says the platform has already revealed regulatory hot spots. “We’ve found changes of function in proteins in sites that are not the classic active sites of these proteins—and these are proteins that have been screened by dozens of companies in the past.”
Meanwhile, Black Diamond Therapeutics, which since December has raised over $100 million, is taking a wildly different approach to allostery. Like many others in the field, the company is exploring how conformational changes affect a protein’s function. But instead of drugging those sites, Black Diamond is looking for allosteric mutations that cause a conformational shift, locking a protein’s active site into the “on” position.
Although researchers were aware of allosteric mutations, they were not well understood. “What we’re saying is that allosteric mutations—just like compounds binding at allosteric sites—cause a change in the protein,” and that can drive cancer, Black Diamond CSO Alex Mayweg says.
Even though the mutations occur away from the active site, Mayweg says, they change its shape—so much so that Black Diamond researchers are convinced they can design inhibitors that spare healthy proteins. Moreover, the altered active site looks similar across several proteins that feature allosteric mutations, meaning one compound could potentially address a swath of cancers.
Under lock and key
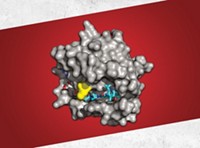
Despite the investment flooding into allostery, biotech executives stress that the field still has to overcome some scientific challenges. For one, technical advances are dramatically improving researchers’ ability to see proteins in their natural state, but getting reliable images remains difficult. Because industry spent so many years working on active-site inhibitors, most of the publicly available protein structures are fragments of the active site.
“One of our big focuses at Relay has actually been reviving the lost art of making full-length proteins and then being able to do structural studies of a full-length protein, frequently in complex with regulatory subunits,” says Don Bergstrom, the firm’s head of R&D.
Advertisement
And even if researchers manage to capture a good image of their protein target, they are feeding it into computational models that are not for the faint of heart. To understand the conformational differences between healthy and mutant proteins, Relay runs simulations of million-atom systems in which the movement of a single atom can impact its surroundings.
Another issue is that the giant compound libraries that companies amassed over the years are full of molecules designed to bind to the active sites of kinases or have a high affinity for G protein–coupled receptors. They are not very useful in the hunt for allosteric modulators. “Allosteric sites are just fundamentally different,” Vanderbilt’s Lindsley says.
HotSpot’s Montagu agrees that developing the right chemical matter will be key. The biotech firm evaluated structures found in chemical libraries for potential activity against allosteric sites and found that only a tiny fraction of the compounds were a good fit. Its scientists took that portion and are now assembling their own library that is better suited to nature’s regulatory pockets.
Others warn that identifying a promising target is only the first step toward new therapies. “All of the approaches are very clever and elegant, but at the end of the day, you’ve still got to do drug discovery, and drug discovery is difficult,” Bicycle’s Keen cautions. “However powerful your computational methods are, however powerful the biology, you’ve still got to convert those molecules into medicines. And that’s a lot of work.”


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter