Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Computational Chemistry
Exploring chemical space: Can AI take us where no human has gone before?
Artificial intelligence is helping us find novel, useful molecules. For the field to really take off, though, these tools will need to be accessible to the wider chemistry community
by Sam Lemonick
April 6, 2020
| A version of this story appeared in
Volume 98, Issue 13

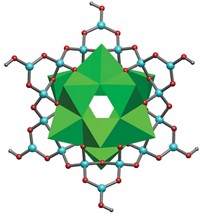
Credit: Jean-Louis Reymond
What’s out there? It’s a question humans have been asking for as long as there have been humans. Our ancestors struck out across continents and oceans to make the unknown known. Now, we’re using telescopes and robots to explore what the universe is like beyond our solar system.
In brief
Chemical space contains every possible chemical compound. It includes every drug and material we know and every one we’ll find in the future. It’s practically infinite and can be frustratingly complex. That’s why some chemists are turning to artificial intelligence: AI can explore chemical space faster than humans, and it might be able to find molecules that would elude even expert scientists. But as researchers work to build and refine these AI tools, many questions still remain about how AI can best help search chemical space and when AI will be able to assist the wider chemistry community.
Outer space isn’t the only frontier curious humans are investigating. Chemical space is the conceptual territory inhabited by all possible compounds. It’s where scientists have found every known medicine and material, and it’s where we’ll find the next treatment for cancer and the next light-absorbing substance for solar cells.
But searching chemical space is far from trivial. For one thing, it might as well be infinite. An upper estimate says it contains 10180 compounds, more than twice the magnitude of the number of atoms in the universe. To put that figure in context, the CAS database—one of the world’s largest—currently contains about 108 known organic and inorganic substances, and scientists have synthesized only a fraction of those in the lab. (CAS is a division of the American Chemical Society, which publishes C&EN.) So we’ve barely seen past our own front doorstep into chemical space.
The sheer size of chemical space can make it frustrating for explorers. Finding useful molecules for a given application can mean navigating dozens of dimensions simultaneously. An effective drug molecule, for instance, might be found where ideal size, shape, polarity, solubility, toxicity, and other parameters intersect.
Right now such a search takes considerable time and expertise. Every molecule that we know of today has “consumed a PhD or is an entire paper,” says Anatole von Lilienfeld, a physical chemist at the University of Basel.

The size and complexity of chemical space make computational tools attractive for exploring it, and the computational tool du jour is artificial intelligence. AI encompasses a variety of techniques, including deep learning and decision trees. These methods are promising, not just for their speed—an AI algorithm can race through thousands of molecules in the time it would take a human to consider one—but also for their ability to consider dozens of dimensions at once. And because AI models can be built with little prior knowledge of the rules or practices of chemistry, these techniques may also be able to find areas of chemical space that chemists’ biases prevent them from searching.
Chemists around the world are turning to AI to help them find new medicines, materials, or consumer products. But experts are still arguing over the best ways to use AI. Should an AI explorer be allowed to roam freely in the chemical wilderness? Or should chemists hold their algorithms on a tight leash, confining them to the area of chemical space where humans think promising molecules will be found? Should AI algorithms learn on their own, or do they need to be taught the physical laws of the universe to search effectively? And how can human chemists and AI work together effectively?
If scientists can answer those and other questions, the result might be AI algorithms and workflows that almost any chemist can use to search chemical space. Off-the-shelf AI tools could be similar to quantum chemical software like Gaussian that offer chemists everywhere powerful computational abilities. Once we reach that point, AI will be able to travel rapidly into the dark areas of chemical space, where no human has gone before.
Search engines
Few chemists are exploring chemical space for exploration’s sake alone—although Jean-Louis Reymond of the University of Bern might seem to fit that description. Reymond has built a succession of databases containing possible molecules that can be made from a certain number of atoms. For instance, his latest database, called GDB17, holds more than 166 billion organic molecules, each made from 17 atoms of only carbon, nitrogen, oxygen, sulfur, and halogens (hydrogens are not counted). When he started making databases like these, he says, “it seemed like nobody had bothered to ask, ‘If we do everything, what do we have?’ ”
By the numbers
10180
An upper estimate of the number of possible molecules
1080
Estimated number of atoms in the universe
1060
An estimate of the number of possible small organic molecules
108
The number of organic and inorganic substances in the CAS database
Sources: Current Topics in Medicinal Chemistry 2006, DOI: 10.2174/156802606775193310; The Mathematical Gazette 1945, DOI: 10.2307/3609461; Acc. Chem. Res. 2015, DOI: 10.1021/ar500432k; Todd Willis interview.
That question is important, Reymond says, because it can lead to useful combinations that no one has considered before. Reymond is looking for possible drug molecules and other useful compounds. And although when he began his explorations in 2005, he didn’t use AI, Reymond says he uses it now to guide his enumeration. Rather than predict every possible atomic combination, he streamlines his efforts by using AI to predict what compounds might be drugs and what molecules can be successfully synthesized. “We have to do that because we can’t use the exhaustive approach,” he says. The databases would be too large to fully enumerate.
AI algorithms vary, but the basic promise of the technology is that with the right kind and quantity of training data, they can learn to understand patterns and then recognize them in unfamiliar data. Image recognition is an area where AI has excelled. For example, algorithms today can be trained on millions of different images, with some guidance or reward system that helps them identify, say, which of the images contain dogs. With enough training, an algorithm can reliably analyze any image and predict whether it contains a dog.
But training an AI algorithm to find a molecule is not quite the same. For one thing, a picture of a dog is a 2-D set of pixels whose color and shading can vary along a continuum. A molecule is a set of discrete atoms at specific distances in three dimensions. Put one pixel out of place, and you can still identify the dog. Put one atom out of place, and you likely change the molecule’s properties.
Another important difference between image recognition and molecule prediction is data availability. The internet gives computer scientists access to billions of images with which to train their models, but chemical data can be comparatively sparse. A data set of melting points compiled by the late Drexel University computational chemist Jean-Claude Bradley has about 29,000 entries. That’s a big number for chemistry, but an AI algorithm tasked with exploring millions or billions of molecular possibilities might need more training data to make accurate predictions across that whole space.
Still, the melting point data are freely available online, which is more than can be said of a lot of the world’s chemical data. Chemical and pharmaceutical companies typically keep their data private. And databases like the one compiled by CAS often require users to pay for access. Although free databases like PubChem exist, some chemists think their information can be less reliable.
Chemists will need to change the way they think about data if AI is going to get better at searching chemical space, says Anne Fischer, who manages several programs in this area for the US Defense Advanced Research Projects Agency. Training AI algorithms on negative data—as in, this photo does not contain a dog—helps make them more accurate. Chemists aren’t reporting negative results, Fischer says.
Pierre Baldi, a computer scientist at the University of California, Irvine, thinks data availability is key to enabling AI exploration of chemical space. “Discovery of drugs and things that benefit all of us could be accelerated by making data more readily open and available within the broad chemical sciences community,” he says. He argues that organizations that own chemical data without making them openly available impede the application of AI across chemical space. For example, Baldi points to ACS, which owns data in the journal articles it publishes and in databases like those compiled by CAS.

Susan R. Morrissey, director of communications for ACS, says the organization feels it’s important to ensure its data are used properly and adds that ACS journals welcome requests to analyze text and data to advance science. “In these cases, we are working closely with the individuals concerned to understand their needs and goals and to ensure the appropriate license and access rights are provided,” she says.
If sufficient, meaningful data aren’t available, the alternative is to code the rules of chemical space into an algorithm. That practice is a bit controversial among chemical space explorers. For instance, image-recognition AI algorithms are typically given few rules or none at all. Computer scientists show the algorithm a picture of a dog, but they don’t explicitly tell it that a dog has two ears, two eyes, a tail, and so forth.
More data are available for some areas of chemical space, like the regions inhabited by small-molecule drugs, because chemists have spent longer exploring them or because there was more incentive to investigate them. In those places, experts say, it may make sense to encode fewer, looser rules, to let the algorithm learn for itself what patterns in molecules impart various properties. This kind of unsupervised learning can yield better results than having lots of rules because it is less biased by human intervention. It can also be a useful approach when the rules are complex or not well understood, like what makes a drug candidate toxic.
But in parts of chemical space with sparse data, experts might need to be more hands on. When good data sets contain only hundreds of points, algorithms can learn faster if they don’t have to teach themselves fundamental or empirical rules of nature, says Ghanshyam Pilania, a materials scientist at Los Alamos National Laboratory. The trade-off is that encoding these rules inevitably means encoding human bias about where useful molecules will be found. “The potential for discovery gets limited” if you tell the AI algorithm too much, Pilania says. He’s used AI algorithms to search for new materials for radiation detectors and electronics.
Pilania and others have found that both approaches have their place in exploration. “At different stages we might want to cast the net wide,” with unsupervised learning, according to Nathan Brown, head of cheminformatics at the drug discovery and development firm BenevolentAI. At other stages, the search may need to be more constrained, especially as Brown’s group narrows in on candidate molecules.
Deciding on which approach to use comes down to how much you think your own expertise biases your search, says Matthew J. McBride, who helps manage a CAS service that uses AI to help customers navigate the intellectual property space of molecules—meaning what compounds are patented and what areas of chemical space might be unclaimed.
McBride has faith that AI will surprise scientists by finding things they didn’t expect. Technology has often surprised humans like that, revealing uses outside the original design, he says.
Molecule generator
Whether AI can or should find those surprises is somewhat contentious within the chemical space exploration community. That’s because community members have strong feelings about data interpolation—finding new data points within a defined set of possibilities—versus extrapolation—finding data points outside the known range. Asking AI to extrapolate is risky and should be done with caution, says Clémence Corminboeuf of the Swiss Federal Institute of Technology, Lausanne (EPFL), who has used machine learning to search for new catalysts.
Researchers have demonstrated some successful AI extrapolation in computer science. But because AI algorithms learn from the training data sets that scientists build and provide them, experts are generally suspicious about algorithms touted as being able to make reliable predictions outside the scope of those data sets.
Advertisement
This isn’t to say that interpolative methods can’t find new molecules. For example, the University of Basel’s von Lilienfeld with Rickard Armiento of Linköping University used AI to search a space of about 2 million compounds, all having a common crystal structure, and found a stable structure whose aluminum atom is in a negative oxidation state, something that had been reported only once before (Phys. Rev. Lett. 2016, DOI: 10.1103/PhysRevLett.117.135502).
In that research, AI was analyzing molecules, but it wasn’t proposing new ones. The group hit on this rare compound’s structure by swapping main-group element atoms into the crystal’s basic architecture to create a library of all possible variants.
But some chemists are using AI to find totally new molecular structures. The so-called generative algorithms they’re using to do that can draw never-before-seen compounds using the AI’s understanding of how molecular composition and structure impart properties. This method reverses the way many groups use AI to search chemical space, which is by interpreting properties from databases of known structures.

Koji Tsuda of the University of Tokyo is among the generative algorithm vanguard. In a 2018 paper, his group demonstrated an AI method for inventing molecular structures that absorb light at certain wavelengths (ACS Cent. Sci., DOI: 10.1021/acscentsci.8b00213). What was interesting, Tsuda says, was that the group’s AI algorithm did not manipulate molecules’ aromaticity to control what wavelength each outputted compound absorbed, a strategy that humans often use. The AI instead manipulated the transitions between nonbonding and antibonding orbitals in each compound to achieve the desired outcome. Tsuda says that’s likely because his group trained the model on a readily available database of small-molecule drugs rather than molecules with large aromatic systems. The AI algorithm wasn’t privy to the kinds of aromaticity that a human chemist might use to solve this problem.
Generative models have been making headlines in drug discovery in the past several months. Insilico Medicine announced in September that one of its models had generated candidate kinase inhibitors for treating fibrosis (Nat. Biotechnol. 2019, DOI: 10.1038/s41587-019-0224-x); in January, it showed that another model generated possible treatments for COVID-19 (bioRxiv 2020, DOI: 10.1101/2020.01.26.919985).

The company’s work has met with some skepticism. W. Patrick Walters and Mark Murcko of biotechnology firm Relay Therapeutics published a critique of the September results, pointing out Insilico’s most promising molecule’s structural similarity to known kinase inhibitors that were included in Insilico’s training data (Nat. Biotechnol. 2020, DOI: 10.1038/s41587-020-0418-2). In other words, the molecule that Insilico found wasn’t exactly groundbreaking, according to the critics. “One has to ask whether a paper reporting work in which a team of chemists substituted an isoxazole for an amide carbonyl to generate a compound that is roughly equipotent with published compounds would be reviewed, let alone published,” Walters and Murcko write.
In a response, Insilico CEO Alex Zhavoronkov and the University of Toronto’s Alán Aspuru-Guzik, coauthors on the 2019 paper, defend their work as a proof of concept and argue that despite similarities between their novel compound and known kinase inhibitors, small structural changes can impart large functional differences (Nat. Biotechnol. 2020, DOI: 10.1038/s41587-020-0417-3). They reiterate that their generative AI did as it was asked: it found new, unique kinase inhibitors.
In closing their critique, Walters and Murcko write, “AI methods have moved beyond the specialist realm that they have occupied for the past 30 years,” suggesting that chemistry may have already reached the point where AI exploration tools can be used by almost anyone. In that context, the two call on the drug discovery community to develop guidelines to better contextualize AI results, such as publishing the contents of training sets. The 2019 paper reported which papers, patents, and databases it drew from to train its algorithms but didn’t include the training data sets.
AI takes the wheel
More openness is a common request from AI experts working in chemical space exploration. In one way, it’s a bit of a paradoxical plea. Powerful AI models like deep neural networks are inherently impenetrable; there’s no real way to open the hood and figure out how they make their predictions. But scientists would like AI predictions to come with error bars. “I think uncertainty measures are necessary” for AI methods to become off-the-shelf tools, says the EPFL’s Corminboeuf.
That’s especially true when chemists have to rely on small data sets to train their AI programs, says Los Alamos National Lab’s Pilania. When it’s not possible to get or generate more data, he says, “it becomes very important to understand our confidence in a machine-learning model’s predictions.”
Implicit in Corminboeuf and Pilania’s desire for uncertainty quantification on AI predictions is the idea that a human expert will be evaluating what the model produces. Humans play a role in almost every step of AI chemical space exploration. Humans decide what types of AI algorithms to use, they build the data sets to train their models (a potential source of bias, researchers have shown), they adjust and retrain the models, and then they analyze the results or attempt to synthesize the predicted molecules.
At least for now. Zhavoronkov and others are trying to take humans out of the equation when they can. “Our AI right now can do completely driverless” exploration, he says of Insilico’s models, meaning that after being given some parameters, the algorithms can computationally generate and screen molecules that are likely to fit the user’s needs.
Most chemists using AI to explore chemical space want to find ways to let computers take over human tasks, but that’s different from replacing human chemists. Jason Hein, who uses AI to understand reaction mechanisms at the University of British Columbia, and others say they think AI tools will empower human chemists. “When I joined grad school, I joined to cure cancer,” Hein says. But 3 years into his PhD he was trying to build a tool to better separate spots on a chromatography instrument’s readout. Hein—and many others who spoke to C&EN—think that AI will alleviate some of the drudgery of lab work, allowing researchers and graduate students to devote more creative energy to solving big problems.

The question is, When will AI be ready for that? Or is it ready right now? Once again, it depends on whom you ask. Connor Coley of the Massachusetts Institute of Technology and Broad Institute of MIT and Harvard uses AI to predict whether molecules found in chemical space can be synthesized. He thinks we’re getting close. He sees most of the AI chemical space explorations going on now as proofs of concept, but he thinks they will help show chemists what approaches work best for different problems. “I think we’re just waiting for these algorithms and workflows to be commoditized,” he says.
Still, a one-size-fits-all AI approach to searching chemical space isn’t likely to ever emerge. One reason is that AI right now struggles to make predictions for large regions of chemical space. “If you explore very large chemical space, as you move [across it], physical mechanisms are changing,” Pilania says. For instance, high- and low-temperature superconductors work in different ways, and AI methods often can’t deal with the rules changing like that.
Some chemists feel as if they can see their AI methods just starting to mature. Two years ago, Heather Kulik of MIT reported an AI method for finding inorganic compounds that can act as switches and sensors. But now she calls the space her team explored then “kind of a toy problem.” In her group’s latest paper, the scientists explored a region of chemical space in which they didn’t have enough data to train their model in a satisfying way. But their active-learning AI models, which can learn as they search, were still able to narrow that space down to its most promising regions and find redox complexes that could be used in batteries (ACS Cent. Sci. 2020, DOI: 10.1021/acscentsci.0c00026).
And others think AI is ready now. Companies like Insilico and BenevolentAI believe that they can already discover useful drugs with AI. And chemists like Tim Cernak of the University of Michigan think off-the-shelf AI tools for chemical space exploration, though very new, are already available for anyone to use.
Cernak points to products like Alphabet’s TensorFlow, cloud-based software that anyone can use to perform machine-learning tasks. Although TensorFlow isn’t made specifically for exploring chemical space, Cernak says users can upload their data and objectives, and the software will attempt to solve a problem using different AI approaches. In the same way a chemist might grab bottles of various metal salts off the shelf to try in a reaction, TensorFlow will try out these methods to find the best one for the job, he says. Whether the average chemist is comfortable using a tool like TensorFlow, however, is another question. And Cernak notes that AI isn’t good enough yet to make experimental verification obsolete.
AI algorithms have already demonstrated that they can recognize—and maybe invent—compounds that not only follow the laws of physics but also have properties that humans are looking for. They are fast and tireless and can be adapted to a variety of tasks. In many ways they seem perfectly suited to the job of exploring chemical space, like the probes and rovers that are exploring parts of our solar system that remain beyond human reach. What remains to be seen is how long these powerful tools will remain in the hands of specialists and what we might find when more chemists can use them effectively.
CORRECTION
This article was updated on April 7, 2020, to correct the description of a crystal containing an aluminum atom in a negative oxidation state. It was referred to incorrectly as a molecule.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter