Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Probing Large Protein Systems
Approach yields first detailed analysis of 456-protein nuclear pore complex
by Stu Borman
December 3, 2007
| A version of this story appeared in
Volume 85, Issue 49
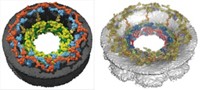
The unnamed technique combines experiment with computation to calculate the structures of large biomolecular complexes. The developers of the technique—NPC specialist Michael P. Rout and mass spectrometrist Brian T. Chait of Rockefeller University; computational structural biologist Andrej Sali of the University of California, San Francisco; and their coworkers-used the approach to obtain the structure of the yeast NPC, which is a massive, membrane-embedded assembly that controls biomolecular traffic between the nucleus and cytoplasm (Nature 2007, 450, 683 and 695).
Crystallography and NMR spectrometry can reveal the structures of proteins and biomolecular assemblies, but not those as large and flexible as NPCs. Electron microscopy (EM) can be used to view large complexes like NPCs, but it cannot distinguish the relative positions of proteins in these assemblies. The new technique makes it possible to determine the architecture of very large complexes and to localize their molecular parts.
"We can resolve the relative positions and arrangements of each and every one of the components of the NPC," Rout says.
"For the first time, we know the positions of the 456 proteins in the NPC," Sali adds. "Previously, we knew little more than the overall shape."
The resolution of the new NPC structure is much lower than the atomic-level resolution achievable on smaller structures with crystallography and NMR. However, "with time, we should be able to collect additional data and improve on our computation to improve the resolution," Sali says. High-resolution crystal and NMR structures of NPC component proteins can now be fitted into the overall structure and used to help build a higher resolution view, he notes.
The technique draws upon data from different types of experiments "as restraints for an optimization method that explores the possible configurations of proteins in the complex," explains John D. Aitchison of the Institute for Systems Biology, Seattle. "The computer explores the possible fits" among components, as if they were puzzle pieces. "If you do this thousands of times and average the solutions, the picture that emerges will be the correct one," Aitchison says.
"Maybe the surprise is that the kind of proteomic and biophysical data we used, which is normally not considered explicitly structural in nature, could yield such an informative structure," Chait says.
The new structural portrait reveals "how the NPC might have evolved through a series of duplications and divergences" of its constituent protein types, called nucleoporins, "resulting in the very simple modular architecture we have today," Sali says. It also suggests how related membrane-based protein complexes might have evolved. And it could inspire future experiments on transport through the NPC and how the complex assembles biosynthetically.




Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter