Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Synthesis
Enzyme By Design
Computational modeling and directed evolution work together to jack up the activity of catalytic proteins
by Celia Henry Arnaud
August 19, 2013
| A version of this story appeared in
Volume 91, Issue 33

Natural enzymes have an exquisite ability to selectively and efficiently catalyze biochemical processes. Scientists have a great interest in harnessing that ability and putting these protein catalysts to work in synthetic applications. To do so, they have been using computer modeling to design enzymes for target reactions and then synthesizing them in the lab. But those enzymes are falling far short of the talents of natural enzymes.
By instead using the designed enzymes as a starting point and tweaking them further through directed evolution, scientists are seeing more success. In directed evolution, the researcher introduces random mutations into genes that code for the production of the enzymes and then screens the resulting enzymes to determine their new activity. In this way, scientists have been able to boost the catalytic activity of designed enzymes by factors of hundreds, or even thousands.
Those improvements suggest that computational design has come a long way. But they also hint that there are further gains to be had.
Most of the success stories in enzyme design so far have come from David Baker’s lab at the University of Washington, Seattle. Even so, Baker isn’t afraid to acknowledge the current shortcomings in computational protein design.
“If the methods were perfect, our initially designed enzymes would have very high activity, and it wouldn’t be possible to improve them,” Baker says. “When we do computational design, we get in the ballpark, but we don’t get the details completely right. It can mean three orders of magnitude difference in enzymatic activity.”
A striking recent example is the case of retro-aldolase, which catalyzes cleavage of a complex ketone to an aldehyde and acetone. Donald Hilvert at the Swiss Federal Institute of Technology (ETH), Zurich, had previously teamed up with Baker and others to design retro-aldolases from scratch (Science 2008, DOI: 10.1126/science.1152692). Hilvert’s team subsequently improved the enzymes via directed evolution (Protein Sci. 2012, DOI:10.1002/pro.2059).
But the researchers wanted to push the retro-aldolases even further.
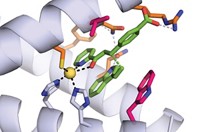
What was surprising is how dramatically the active site changed in the process. In the original designed enzyme, the reaction took place at the intended catalytic lysine in the enzyme’s amino acid chain, which was located in a hydrophobic binding pocket. After eight rounds of directed evolution, catalysis was occurring at another lysine in a different binding pocket. The best of the reported evolved enzymes is more than 4,000 times as active
Several things had to happen to get that new active site, Hilvert says. The first rounds of directed evolution introduced several beneficial mutations, including the new lysine and substitutions that altered one of the turns, or loops, in the enzyme’s structure, creating the new binding pocket for the reactant molecule.
“The new binding pocket where the ligand sits wasn’t present in the original design,” Hilvert says. “The computer couldn’t have predicted its existence because you needed a big conformational change in one of the loops to open up this slot.”
The resulting enzyme is promiscuous, which provided an avenue for evolving the new active site. Both the original and the new lysine residues can catalyze the reaction. In subsequent rounds of directed evolution, an asparagine and a tyrosine were added that help the second lysine outcompete the first one for substrate.
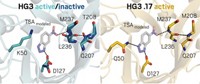
In another example, Dan S. Tawfik at Weizmann Institute of Science, in Israel, and coworkers also see dramatic improvements when they use designed enzymes in directed evolution. The researchers started with another family of enzymes designed by Baker—Kemp eliminases (Nature 2008, DOI: 10.1038/nature06879). The Kemp elimination is a ring-opening reaction that involves removing a hydrogen atom from a carbon and simultaneous cleavage of a nitrogen-oxygen bond in benzisoxazoles.
Tawfik’s team started by identifying 23 locations outside the active site where amino acid changes might reinforce the structure of the originally designed enzyme without affecting its activity. Because they didn’t know which changes would work best, they added them in a combinatorial fashion in the first two rounds of directed evolution. Ultimately, 10 of the mutations ended up in the evolved enzyme variants.

The researchers then conducted another 16 rounds of directed evolution to optimize the catalytic activity (Proc. Natl. Acad. Sci. USA 2012, DOI: 10.1073/pnas.1121063109). Catalytic improvements plateaued after 13 rounds.
“Early mutations give you large improvements,” Tawfik says. “The later mutations just reinforce the effects of the early ones that change the chemistry and the substrate binding.” Using a different system, Tawfik and coworkers have shown how directed evolution can run up against diminishing returns (Nat. Commun. 2012, DOI: 10.1038/ncomms2246).
In the case of the Kemp eliminases, it’s difficult to understand how specific changes improve the catalytic activity, Tawfik continues. Therefore, it’s hard to know how to use the directed evolution to improve the algorithm for future designs.
“With few exceptions, it’s impossible to figure out on an atomic level exactly how the mutations altered the design in a way that makes the active site more efficient,” he says. “The mutations are in residues that aren’t in direct contact with the substrate.”
Most likely, those mutations slightly reshape the active site and spatially fix the catalytic amino acid residues, Tawfik notes. “In natural enzymes, the catalytic residues would be held in place by a whole network of hydrogen bonds,” he says. “That’s what we think many of the mutations in directed evolution do. It’s just that the hydrogen bond networks are so complex that it’s impossible to say from the structure how they contribute energetically.”
But Hilvert still thinks the up-front design is important because it “gives you a foot in the door.” His team has tried with little success to find good starting points for directed evolution by screening random libraries. Many of the proteins in random libraries have no catalytic activity. Starting with a protein derived from computational design provides at least minimal activity.
One problem with computational design is the difficulty in accurately modeling protein loops. These enzyme structural features are disordered, floppy regions. But in computational design, the loops are usually assumed to be fixed. In the case of the retro-aldolases, crystal structures showed that the loops in the actual enzymes didn’t match the design.
“This simplifying assumption that the protein is rigid is not valid,” Hilvert says. “In a case like ours, with a protein where the active site is constructed from loops of varying length and mobility, that could be a problem.”

But that floppiness is what makes the designed proteins evolvable, Tawfik asserts. Natural evolution usually takes advantage of catalytic residues that can already perform the desired chemistry and repositions the substrate or the catalytic residues to improve the activity. Without at least some activity, no matter how low, “you wouldn’t have a starting point for evolutionary optimization,” he says.
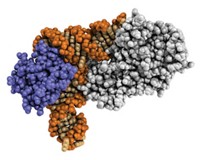
And sometimes one amino acid is all it takes to transform a protein with no catalytic activity into one with just enough activity to be a candidate for directed evolution. That’s what a team led by Ivan V. Korendovych of Syracuse University has done with the calcium-binding protein calmodulin.
“We decided we’re not going to worry about making a precise fit. We’ll just see if the calmodulin scaffold can potentially accommodate the substrate,” Korendovych says. The researchers did some initial calculations to see which places in the calmodulin scaffold were good candidates for inserting catalytic residues. By changing a single phenylalanine to glutamic acid, they found a protein that could catalyze the Kemp elimination (Proc. Natl. Acad. Sci. USA 2011, DOI: 10.1073/pnas.1018191108). Seven rounds of directed evolution later, they had a catalyst that was more than 200 times as active as the starting catalyst (Angew. Chem. Int. Ed. 2013, DOI: 10.1002/anie.201302339).
When it comes to improving enzyme design, one way is to better understand all the dance steps an enzyme has to progress through during catalysis. “There’s still a lot of mystery about how enzymes achieve their high levels of activity,” Baker says. “Think about what an enzyme has to do: It has to bind the substrate with reasonable affinity and selectively stabilize the transition state. Then it’s got to release the product, so it can’t bind the product too tightly. If there are multiple intermediates, it’s got to interact with all of them.” If the enzyme stumbles anywhere along the way, the reaction won’t happen. And the structures at the various steps might differ from one another by less than an angstrom.
To gain some of the understanding needed to design new enzymes, Baker and his coworkers are dividing the problem into smaller slices. As a first step, they are taking catalysis out of the picture and focusing just on how noncatalytic proteins bind small molecules.
“One of the reasons we went back to small-molecule design is we wanted to get control of really precise protein-small molecule interactions,” Baker says. “In an enzyme, there are all these steps. It’s hard to pin that down. In a protein-small molecule binding interaction, it’s a simpler problem.”
Scientists seem destined to continue using computational design and directed evolution in tandem. By plugging information gleaned from directed evolution back into design algorithms, those algorithms will provide even better starting points.
“Computational design is successful in making the path to a catalyst shorter,” Korendovych says. “It’s not quite able to make the leap directly to an efficient catalyst alone, but it’s making the path shorter.”
“The hope is that, in the future, computation will give us much better starting points,” Hilvert adds. “I’m convinced that the better the starting point, the easier it will be to evolve.”
Tawfik agrees. In the three Kemp eliminases his team has evolved, the researchers found that the higher the starting activity, the greater the improvement from directed evolution. “Even small improvements in design can be translated to dramatic improvements in the end point,” he says.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter