Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Pharmaceuticals
Clues for Overcoming HIV Drug Resistance
Researchers are gaining insights into how some anti-HIV drugs avoid resistance mechanisms
by CELIA M. HENRY, C&EN WASHINGTON
May 24, 2004
| A version of this story appeared in
Volume 82, Issue 21
Human immunodeficiency virus evolves rapidly in patients, making the development of drug resistance a major problem in combating the virus. Some drugs are better than others at avoiding resistance mechanisms, but little is known about what gives them this edge.
In two recent papers, Eddy Arnold, a professor in the Center for Advanced Biotechnology & Medicine and the department of chemistry and chemical biology at Rutgers University, Piscataway, N.J., and his colleagues propose how they think some HIV reverse transcriptase (RT) inhibitors manage to successfully evade the effects of resistant mutations. A coauthor on both papers is virologist Stephen H. Hughes of the HIV Drug Resistance Program at the National Cancer Institute (part of the National Institutes of Health). Arnold and Hughes have collaborated on studies of the structure and function of HIV RT since 1987.
"Unless you consider drug resistance, the virus is going to beat you," Arnold says. "We know that HIV is a moving target. We know that a moving target is hard to hit, particularly when its movements are unpredictable. What we've tried to do is consider in the strategy as many resistance mutations as we're aware of."
RT is a viral enzyme that transcribes the virus's single-stranded RNA genome into DNA. Nucleoside RT inhibitors (NRTIs) are incorporated into the viral DNA by the enzyme, blocking growth of the DNA chain. The common AIDS drugs AZT (zidovudine) and 3TC (lamivudine) belong to this class of pharmaceuticals.
Resistant virus strains develop in patients treated with either AZT or 3TC. In the case of 3TC, mutations in RT cause steric hindrance, selectively blocking the incorporation of the inhibitor into the DNA chain without affecting the incorporation of normal DNA building blocks.
The mechanism of AZT resistance is quite different. The resistant viruses still incorporate AZT, temporarily terminating the growth of the DNA chain. But in resistant viruses, AZT is selectively excised by the mutant RT, allowing the viruses to complete the synthesis of viral DNA.
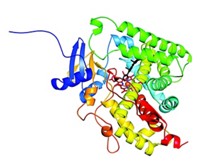
The nucleotide drug tenofovir retains its potency against a number of the viruses containing mutations that cause resistance to other NRTIs. Arnold and his colleagues, including scientists from Rutgers, NIH, and Gilead Sciences, report crystal structures of two RT-DNA complexes before and after tenofovir has been incorporated at the end of the DNA [Nat. Struct. Mol. Biol., 11, 469 (2004)]. The crystal structures determined by lead author Steven Tuske and his teammates reveal the interactions between tenofovir and the HIV RT polymerase active site.
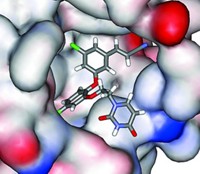
Most important, tenofovir remains completely inside the "substrate envelope," with no protrusions that could provide a handle for drug-resistant mutations. "Reverse transcriptase uses a series of substrates--normal nucleotides--that bind to the enzyme and are used to make viral DNA," Arnold says. "If the drug is smaller than the normal substrates, if it fits within that substrate envelope, you can imagine that the enzyme can't selectively get hold of the drug and block its incorporation without interfering with the incorporation of the normal substrates."
Celia A. Schiffer, an associate professor of biochemistry and molecular pharmacology at the University of Massachusetts Medical School, in Worcester, and her colleagues first used the concept of the substrate envelope with another HIV drug target, the enzyme protease [Structure, 10, 369 (2002)]. She defines the substrate envelope as the intersecting region of the active site pocket in which most of the natural substrates overlap. In structure-based drug design, researchers usually build inhibitors to compete with the substrate without necessarily paying attention to the details of how the natural substrate interacts with the enzyme.
"NOT PAYING ATTENTION to those details leaves the inhibitor susceptible to mutations that could selectively impact the inhibitor over the substrate," Schiffer notes. For her work with HIV protease, Schiffer examined more than 200 crystal structures in the literature that showed the enzyme with various inhibitor complexes, but none looked at how the substrates bound. "If you map the inhibitors onto where the substrates bind, you find that the inhibitors protrude beyond the substrates in a variety of locations," she says. Those locations are where the resistance mutations occur.
Schiffer and her colleagues have discussed the importance of the substrate envelope for one mutation in HIV protease [J. Virol., 77, 1306 (2003)]. But more recent work shows that the concept of keeping drugs within the substrate envelope may be a general mechanism for avoiding drug resistance in any rapidly mutating target. "The implications are that you need to pay attention to how the target interacts with its substrate in order to avoid resistance," she says.
For a second class of anti-HIV drugs that target reverse transcriptase, the nonnucleoside RT inhibitors (non-NRTIs), flexibility of the drug molecule appears to be the key to evading resistance.

Arnold and his colleagues describe the role of conformational and positional adaptability in the design of the non-NRTI etravirine [J. Med. Chem., 47, 2550 (2004)]. A Belgian-American team led by the late Paul A. J. Janssen, including scientists at the Center for Molecular Design, Janssen Pharmaceutica, and Tibotec (all subsidiaries of Johnson & Johnson), as well as NIH and the Arnold lab, incorporated known resistance mutations into the screening process and found that successful compounds were able to inhibit both the wild-type (nonmutant) virus and the drug-resistant mutants. This led to the discovery of a family of diarylpyrimidine compounds, known as DAPY derivatives [Bioorg. Med. Chem. Lett., 11, 2235 (2001)]. The DAPY derivative etravirine is undergoing clinical development as an anti-AIDS drug.
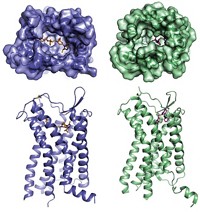
Arnold's colleague Kalyan Das, an associate research professor at Rutgers, proposed that drug flexibility could explain why it was so difficult to get crystal structures of etravirine bound to RT. "A flexible compound may actually bind to the enzyme in a number of different ways. When we grow crystals, the crystals may contain a mixture of different conformations of the enzyme," Arnold explains. "After literally hundreds of tries, Das and [staff researcher] Art Clark were able to cocrystallize etravirine--not by using the wild-type enzyme but by using a clinically important reverse transcriptase mutant. It was only through a very challenging ordeal that we managed to obtain that structure.
"By a process of aggressive screening against different drug-resistant mutants, we've come up with compounds that, when given as single drugs, perform as five-drug combinations in clinical trials," Arnold says. In one-week trials, previously untreated patients experienced a viral load drop similar to that with a five-drug combination [AIDS, 17, 2623 (2003)].
These compounds have conformational flexibility that allows them to bind effectively to resistant RTs. A first-generation drug that targets the same binding site, nevirapine, is rigid and fills the binding pocket. Single mutations cause steric hindrance or lead to loss of favorable contacts with a large, rigid drug like nevirapine. Such mutations can cause "catastrophic resistance" and make the compound "a goner," Arnold says.
"In the case of the DAPY inhibitors, there are conformational degrees of freedom--the drug can assume a different shape that allows the inhibitors to basically wiggle around in the pocket," Arnold says. "Because they're compact, they're also able to jiggle around and reorient and reposition within the binding site." Arnold proposes that the conformational flexibility allows the drug molecules to bind to many different forms of the binding pocket found in drug-resistant RTs. Colleague and study coauthor Paul J. Lewi of the Center for Molecular Design has used computer modeling to study multiple binding modes for the DAPY inhibitors [J. Comput.-Aided Mol. Des., 17, 129 (2003)].
Drug flexibility should be applicable to targets other than RT, according to Arnold. "This should be a fairly general solution to the problem of designing drugs that have to work against targets that vary rapidly--the kinds of targets found in viruses and other infectious agents," he says. "The flexibility of the drug complements the changes in the drug target."




Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter