Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Sialyltransferase Structure
Researchers achieve first structural and mechanistic analysis of enzyme
by Stu Borman
January 26, 2004
| A version of this story appeared in
Volume 82, Issue 4

The first detailed structural and mechanistic analysis of a sialyltransferase, an enzyme that transfers sialic acid groups to cell-surface glycoproteins and glycolipids, is providing insight into how such enzymes work.
Up to now, the membrane-bound nature of sialyltransferases has prevented the enzymes from being purified, expressed, crystallized, and analyzed. Therefore, little has been known about the structures or mechanisms of these enzymes.
That has now changed, thanks to the structure obtained by Natalie C. J. Strynadka and Stephen G. Withers of the University of British Columbia; Warren W. Wakarchuk of the Institute for Biological Sciences of the National Research Council, Ottawa; and coworkers [Nat. Struct. Mol. Biol., published online Jan. 18,<br > http://dx.doi.org/10.1038/nsmb720]. The study also reveals the sialyltransferase's mechanism of action, which could help lead to the development of inhibitors as potential therapeutic agents.
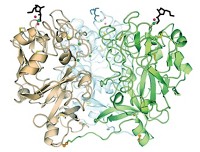
The researchers solved the long-standing problem by removing a small part of the membrane-association domain of a sialyltransferase from the bacterial food pathogen Campylobacter jejuni, making it possible to isolate, crystallize, and analyze the enzyme. Strynadka credits Ph.D. student Cecilia P. C. Chiu for actually purifying and crystallizing the enzyme and determining its structure.
Sialic acid (N-acetylneuraminic acid) groups are found at the ends of many glycans, glycoproteins, and glycolipids, where they play important roles in cellular and molecular recognition and communications. Hence, the structure of the enzyme that transfers these groups to biomolecular acceptors "has long been one of the major goals of structural and mechanistic glycobiology," comments Gideon J. Davies of the University of York, in England.
Sialyltransferases are members of the glycosyltransferase family of enzymes, of which about 20 varieties are known in mammals. The first glycosyltransferase structure was determined in 1994 by Paul S. Freemont of Imperial College, London, and coworkers, and the next was reported in 1999 by Davies' group. "Currently, about 12 distinct structures are known," Davies says, and each has been found to adopt one of two 3-D folds--called GT-A and GT-B. The sialyltransferase analyzed by Strynadka and coworkers has a GT-A fold, but its structure is unique, Davies notes, in that it binds its ligands on opposite sides of the structure from previous GT-A enzymes.
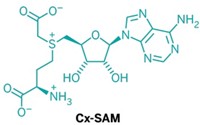
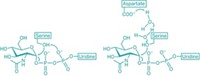
By using a stable analog of cytidine-5'-monophospho-N-acetylneuraminic acid (the enzyme's substrate), Strynadka and coworkers were able to hypothesize a plausible mechanism of action for sialyl transfer. "A histidine acts as a base to assist nucleophilic attack of the acceptor hydroxyl, and a tyrosine is used as an acid to assist the departure of cytidine-5'-monophosphate," explains Chi-Huey Wong of Scripps Research Institute. "Further understanding of the specificity and detailed mechanism of the reaction should provide valuable information for design of new substrates and inhibitors," Wong says.
James C. Paulson, also of Scripps, says the study is an important breakthrough, but he notes the bacterial enzyme has "minimal homology to mammalian enzymes, where most of the interest is." Davies agrees that "it remains to be seen how relevant this structure is to higher organism sialylation, since the Campylobacter enzyme is not closely related, at least at the sequence level, to human sialyltransferases." But Monica M. Palcic of the University of Alberta, Edmonton, says she "would expect mammalian sialyltransferases to exhibit similar topography."


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter