Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Materials
DNA Sorts Carbon Nanotubes
Specific sequences separate nanotubes according to chirality
by Laura Cassiday
July 20, 2009
| A version of this story appeared in
Volume 87, Issue 29

Single-walled carbon nanotubes (SWNTs) show great promise as components of nanoscale electronic devices, but most commercial applications have been stymied by the difficulty in isolating nanotubes of identical chirality from a synthetic mixture. Now, Xiaomin Tu and Ming Zheng of DuPont Central Research & Development, together with Suresh Manohar and Anand Jagota of Lehigh University, have shown that the unique molecular properties of DNA can be exploited to sort SWNTs (Nature 2009, 460, 250).
SWNT synthesis produces a mixture of nanotubes with nonuniform diameters and chiralities and, therefore, heterogeneous physicochemical properties. Having previously shown that a particular DNA sequence could form an ordered structure on SWNTs, Zheng and colleagues reasoned that they might be able to find a DNA sequence to purify each type of SWNT in a synthetic mixture. The problem was identifying the correct DNA molecules among an unfeasibly large number (1018) of possible 30-nucleotide sequences.
To reduce the DNA library to a more manageable size of 350 oligonucleotides, the researchers devised a sequence-pattern-expansion scheme that considered all possible DNA sequences composed of mono-, di-, tri-, and tetranucleotide repeats. They added each DNA oligonucleotide to a random mixture of SWNTs. Then, they used ion-exchange chromatography to separate the 350 solutions into fractions, which they analyzed spectroscopically for the presence of specific DNA-SWNT hybrids.
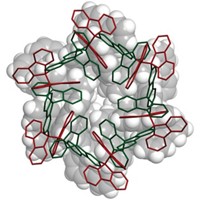
The study yielded more than 20 sequences that could together purify all 12 major chiral semiconducting SWNTs. To explain the SWNT-sorting ability of DNA, the authors propose a model in which the recognition sequences form stable hydrogen-bonded DNA barrels around specific SWNTs. The ordered structure minimizes interactions with the ion-exchange chromatography resin and causes early elution.
"This is a very impressive study that reports the most selective method yet found for isolating specific structural forms of SWNTs from mixed samples," R. Bruce Weisman of Rice University says. "The main limitation is the small scale and high expense." Zheng tells C&EN that the major obstacle to scaling up the method is the high cost of DNA, which could decrease if oligonucleotide suppliers shift their business model to meet the demands of the nanoelectronics industry.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter