Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Proteins In Transition
Chemical Biology: Conformational change proceeds along surprising path
by Sophie L. Rovner
December 17, 2009

Proteins change shape frequently to fulfill their duties. For instance, proteins that organisms use for signaling possess one shape when they're inactive and another shape once they've been activated to transmit a signal. Now, a group of biochemists at Brandeis University have for the first time observed how a signaling protein shifts between these two different conformations.
Conventional wisdom holds that most signaling proteins' conversion from an inactive to an active conformation is triggered by phosphorylation. However, Dorothee Kern and her Brandeis colleagues discovered that a bacterial signaling protein called nitrogen regulatory protein C (NtrC) switches between these two conformations regardless of whether the protein is phosphorylated. With the help of nuclear magnetic resonance dynamics experiments and computational techniques, they found that phosphorylation of the protein occurs only after the conformational change. Phosphorylation stabilizes the active state, which as a result becomes the energetically favored conformation (Cell 2009, 139, 1109).
After the protein carries out its signaling task, it is quickly dephosphorylated. The protein then spends most of its time in the inactive conformation, which is energetically favored when the protein lacks the phosphate group.
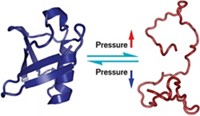
The Brandeis team also tackled another long-held belief, which posits that a protein has to partially unfold to convert from one conformation to another. Kern never bought into that theory. She reasoned that unfolding would be risky because a partially unfolded protein would be vulnerable to degradation or aggregation, which can lead to disease.
Now, Kern's team has demonstrated that NtrC changes conformation without unfolding. The group discovered that nonnative hydrogen bonds form fleetingly within the protein, helping it change shape in a rapid, step-by-step manner in which bonds are broken and formed one at a time rather than all at once. This controlled reconfiguration avoids the risks of unfolding and also lowers the energy barrier that separates the inactive and active conformations.
Kern says NtrC makes a good subject because the molecule is small enough to study with NMR and because it undergoes a large and complex conformational change. She believes the NtrC findings will extend to other proteins. She also expects the insight the research provides into the dynamic way in which proteins work will improve the design of artificial enzymes to carry out desired functions.
The Brandeis team's "studies are remarkable," David D. Boehr, a Pennsylvania State University chemist who studies protein dynamics, notes in a Cell commentary about the work. "They not only reveal new insights into the activation mechanism of NtrC, but they also provide a methodology to elucidate the connections between the energy landscapes of folding and function in other proteins."


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter