Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Pharmaceuticals
Antibacterial Boom And Bust
After a flurry of discovery in the 20th century, the antibacterial drug pipeline is now drying up
by Michael Torrice
September 9, 2013
| A version of this story appeared in
Volume 91, Issue 36

When David L. Pompliano joined GlaxoSmithKline in 2002 as head of biology for infectious diseases, the pharmaceutical company was ending a largely unsuccessful multiyear screening program to search for novel antibacterial drugs.
The program started in the mid-1990s after the publication of the first bacterial genomes. GSK and other pharmaceutical companies thought these genomes held a trove of new targets for antibacterial drugs. The company examined 300 genes, ran 70 high-throughput screening campaigns, and screened between 300,000 and 500,000 compounds per campaign. After seven years, company scientists found just five drug leads. The success rate was about 20–25% lower than rates for other therapeutic areas in the company (Nat. Rev. Drug Discovery 2007, DOI: 10.1038/nrd2201).
Looking back on the vast effort and resources spent on the program, the strategy “certainly did not live up to the promise that everyone was proposing,” Pompliano says.
The GSK story exemplifies the difficulties of discovering new antibacterial drugs. Over the past 90 years, antibacterial discovery has gone from boom to bust. For about three decades in the middle of the 20th century, pharmaceutical companies regularly churned out new classes of the drugs, many of which doctors still use today, such as the penicillins and tetracyclines. But by the 1980s, discovery slowed and companies started leaving the field, drawn by the rise of profitable drugs in other therapeutic areas. As a result, only one successful new class of antibacterial drugs—the antituberculosis drug bedaquiline—has been discovered since the late 1980s.
With the growing numbers of bacterial strains resistant to existing drugs, pharmaceutical experts wonder how to refill the antibacterial pipeline.
The success of early antibiotics saved many lives. In a 1951 story, for example, C&EN reported that, thanks to drug treatment, the pneumonia mortality rate dropped by 50% in the previous decade (C&EN, March 26, 1951, page 1190). And without the ability to manage bacterial infections, successful anticancer therapies that suppress the immune system wouldn’t be possible.
The antibacterial development boom started after the discovery in 1943 of streptomycin, the first antibiotic to treat tuberculosis. Albert Schatz, a graduate student at Rutgers University, and his supervisor, microbiologist Selman A. Waksman, found the compound in Streptomyces bacteria and learned that it killed other microbes. Streptomyces live in soil, and soon pharmaceutical companies across the U.S., Europe, and Japan started similar screens of soil microbes.
These companies were tapping into a microbial war in the dirt that had been going on for millennia, says Richard J. White, a pharmaceutical scientist who worked on antibacterial discovery for 28 years at several companies. The screens pulled out natural products that bacteria produced to slow or kill their microbial competitors.
ANTIBACTERIAL TIMELINE
Number of successful antibacterial
classes discovered per decade
In a typical natural products screening program, company microbiologists would amass soil samples from across the globe. When company employees went on vacations to exotic spots, the microbiologists would send them off with soil sample bags in hand, hoping the employees would snag a rare species that produced a novel compound, White says. Once soil samples came in, the microbiologists would first isolate the multitude of different microbes present and then grow them separately in liquid cultures. The researchers then tested the resulting broths to see whether they could stop the growth of their pathogen of choice, maybe Staphylococcus aureus or Escherichia coli. If they did, the real work started: isolation of the active molecule from the bacterial soup.
From the 1940s through the 1960s, pharmaceutical companies improved this general method and discovered about 20 major antibacterial classes, including the tetracyclines, the macrolides, and the glycopeptide vancomycin.
As scientists studied the major classes, they found out how the antibacterials work. The β-lactams, such as penicillin and cephalosporin, inhibit cell wall synthesis. Tetracyclines, macrolides, and aminoglycosides gum up protein synthesis. And the quinolones disrupt DNA replication.
Medicinal chemists played a significant role in the boom by developing new analogs of the antibacterial classes with improved properties, says Steven J. Brickner, a medicinal chemist who worked at Upjohn and Pfizer. Beecham Research Laboratories, an English company that through mergers became part of GSK, produced several important derivatives of penicillin. Its medicinal chemists changed penicillin’s side chains to help the drugs overcome resistant microbes and to attack a wider range of bacterial species. For instance, methicillin, developed in 1959, has a bulky 2,6-dimethoxyphenyl side chain, which shields the compound from some β-lactamases, the enzymes that enable bacteria to resist penicillin and other β-lactam drugs.
Discoveries of new classes started to taper off during the 1970s. Companies started to see diminishing returns in their natural products screens, as they exhausted most of the low-hanging fruit.
As the slowdown started, Brickner remembers, researchers would go through liters of black soups to isolate milligrams of a compound—only to realize later it was either something already known or a close derivative of a known antibiotic. In the 1950s, companies had to screen through about 1,000 bacterial cultures to find a compound no one had seen before, says Lynn Silver, who worked at Merck & Co. for more than 20 years. To find daptomycin, which was discovered in 1987 and is one of the last truly new classes to reach the market, scientists had to pick through about 10 million cultures.
In the late 1990s, the industry tried to invigorate antibacterial discovery by turning to genomics. After the publication in 1995 of the genome of Haemophilus influenzae, companies such as GSK thought they could find new drug leads by searching for genes essential for bacterial survival in multiple species. Then, using in vitro assays, they screened for compounds that inhibit the activity of the proteins coded by those genes. The assays enabled high-throughput screens using libraries of synthetic molecules.
The strategy failed for multiple reasons. First, it focused too much on single targets, Silver says. Most successful antibacterial drugs hit more than one target in bacteria: β-Lactams hit multiple proteins involved in cell wall synthesis, and quinolones knock out a couple of topoisomerases, enzymes that help bacteria replicate their DNA. If a compound shuts down just one target, bacteria can easily mutate that gene and rapidly gain resistance to the drug.
The other problem was that chemists synthesized the libraries on the basis of rules for making drugs that hit targets inside and outside human cells. Those rules are not helpful in attacking bacteria. Unlike human cells, for instance, gram-negative microbes have dual-membrane barriers and protein complexes that actively pump out unwanted chemicals. Scientists currently lack an understanding of what allows compounds to slip past all of these barriers.
With the failure of high-throughput screening, many companies shifted their focus to more lucrative therapeutic areas, such as chronic diseases, which must be treated over the patient’s lifetime, unlike bacterial infections. Meanwhile, many still-effective drugs went off patent, filling the market with cheap generics.
At the start of the 1980s, more than 35 major U.S. and European companies were working on antibacterial drugs, says Karen Bush, a biochemist at Indiana University, Bloomington, who worked in the pharmaceutical industry for more than 30 years. Now there are fewer than 10. The exodus has led to a steady decline of new drugs hitting the market.
According to the Infectious Diseases Society of America (IDSA), 16 new antibacterial drugs were approved in the U.S. between 1983 and 1987. In the past five years, just two have been approved (Clin. Infect. Dis. 2013, DOI:10.1093/cid/cit152).
While the pipeline has been drying up, pathogens have been developing resistance to existing drugs, says Manos Perros, head of the infection innovative medicines unit at AstraZeneca. For example, earlier this year, the Centers for Disease Control & Prevention issued a warning about gram-negative pathogens called carbapenem-resistant Enterobacteriaceae (CRE). Carbapenems are β-lactam drugs often used as a last resort in serious infections. One of every two patients with CRE in the bloodstream dies.
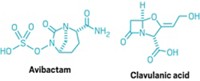
Antibacterial experts see a few strategies to attack resistance. One that has been around since the mid-1970s is combining a β-lactam drug with a β-lactamase inhibitor (BLI) to block the effects of resistance genes. For example, the successful drug Augmentin pairs amoxicillin with the BLI clavulanic acid. Four β-lactam/BLI combination drug candidates are currently in Phase II or III clinical trials.
Other potential tools against resistance are rapid diagnostic tests. If doctors can quickly identify the infectious agent, they could be more selective about which drugs to use. Selective drug use could slow the development of resistance.
Many experts think the future of new antibacterial treatments will depend on luring pharmaceutical and biotech companies into the field. Groups such as IDSA recommend a combination of economic incentives and regulatory reforms to make antibacterial development more financially attractive to companies and investors. Potential incentives could be tax credits for research and development or provisions to extend the exclusivity period for drugs to treat serious infections.
If companies return to the antibacterial field, the resistance problem is surmountable, experts say. “It’s doom and gloom right now, because the pipeline is so empty,” Wright says. “But the reality is we can solve these problems, it just won’t be easy.”
MORE ON THIS STORY
Introduction: Nine For Ninety | PDF
Chemical Connections | PDF
Antibacterial Boom And Bust | PDF
Small Science, Big Future | PDF
Understanding The Workings Of Life | PDF
Chemistry By The Numbers | PDF
Plastic Planet | PDF
The Catalysis Chronicles | PDF
Giving Chemists A Helping Hand | PDF
It's Not Easy Being Green | PDF
Readers' Favorite Stories | PDF
Nine Decades Of The Central Science | PDF
How Chemistry Changed the World | C&EN's 90th Anniversary Poster Timeline
Trying To Explain A Bond
A Toast To C&EN At 90



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter