Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Materials
Protein Stores Copper For Methane-Digesting Bacteria
Structural Biology: Tetramer of four-helix bundles holds a total of 52 copper ions
by Celia Henry Arnaud
August 26, 2015

Engineered methanotrophic bacteria could help us cut atmospheric levels of methane, a greenhouse gas. But scientists first need to understand how the microbes handle the copper required for their methane-oxidizing enzymes.
Scientists have known how copper gets into the bacteria, but they haven’t known how the bacteria store the copper they take up. A newly identified protein is a good candidate for that storage space.
Christopher Dennison of Newcastle University, in England, and coworkers discovered a new protein, called Csp1, that can bind up to 52 Cu(I) ions (Nature 2015, DOI: 10.1038/nature14854). Using bioinformatics, the researchers discovered two other copper storage proteins belonging to the same family in the same bacterium.
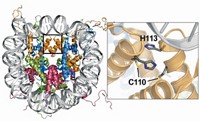
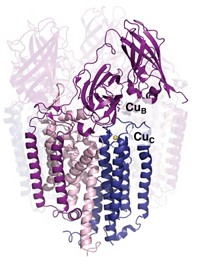
The X-ray crystal structure and in vitro studies of Csp1 show that the tetrameric protein binds 13 Cu(I) ions in each of its four four-helix bundles. The ions line up down the middle of each bundle bound by cysteine residues that point into the bundle’s core.
“This report shows that Cu(I) ‘copper sponges’ do indeed exist and provides atomic-level details on how they might function in copper homeostasis,” says Amy C. Rosenzweig, an expert on metalloproteins at Northwestern University.
And such copper sponges might not be limited to methanotrophs. Additional bioinformatics suggests that many other bacteria have similar copper storage proteins.
“There is a broadly accepted idea in the community that bacteria have very little in the way of an intracellular copper requirement,” says David P. Giedroc, an expert on bacterial transition metal homeostasis at Indiana University. “This paper challenges that view.”
The findings also could help bioengineers working with methane monooxygenases (MMOs), says Ramon Gonzalez, who studies metabolic engineering at Rice University. “It has been pretty much impossible to recombinantly express MMOs in a foreign host,” he says. Adding Csp1 to these strains could pave the way for industrial microbes that use methane as a chemical feedstock, Gonzalez says.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter