Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Environment
Charge-free protein retains its structure and function
A protein that had its charged amino acids swapped for neutral ones remained stable, soluble, and functional
by Erika Gebel Berg
July 20, 2016

Though charge is one of a handful of fundamental factors that dictate a protein’s activity, scientists don’t fully understand how interactions between positive and negative amino acids affect a protein’s structure and function. Now, in a surprising twist, researchers have produced a mutant protein domain that lacks charged amino acid residues, but retains its folding and cellulose-binding abilities. The findings may have implications for protein design, helping scientists build better algorithms for designing proteins with new functions (Biochemistry 2016, DOI: 10.1021/acs.biochem.6b00269).
Charged amino acids are generally accepted as necessary for proper protein folding, solubility, and functions such catalysis, says Jakob R. Winther of the University of Copenhagen. Five out of the 20 amino acids commonly found in proteins are either positively or negatively charged under physiological conditions, and all known soluble proteins have at least a few of these residues. “Predicting charge interactions is incredibly difficult,” Winther says. “One charge will affect another and another. Everything gets tangled in a web of mutual attraction that blurs what’s going on.” To help unravel this web, Winther’s team attempted to create a protein with no charges.
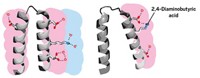
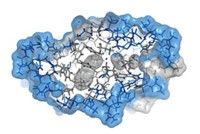
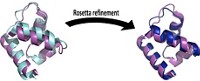
As a first step, the researchers went to the Protein Data Bank—a repository of 3-D structures of large biomolecules—to find the least charged protein or protein domain of 100 residues or more. That turned out to be a cellulose-binding domain from the bacterium Cellulomonas fimi, which contains four charged residues. Next, the researchers used a program to help guide which mutations to make, looking for swaps with neutral amino acids that would produce the most stable protein possible. The winning mutant replaced a lysine, aspartic acid, arginine, and histidine with a methionine, glutamine, methionine, and tryptophan, respectively.
The researchers then expressed their uncharged mutant protein in Escherichia coli and put it through the wringer. They tested its solubility by checking how easily it precipitated in solution and its stability by monitoring how easily it unfolded when mixed with guanidine hydrochloride, which is commonly used to denature proteins. They also determined the protein’s structure with circular dichroism and nuclear magnetic resonance spectroscopy.
Based on all these measures, the uncharged protein seemed largely unchanged from the charged version. Plus, it’s actually more stable than the original protein. “Most proteins unfold with acid or base,” Winther says, because extreme pHs tend to add or subtract a charge from negatively or positively charged residues. As a final test, the researchers ran an assay to see whether the protein still binds cellulose. “To our surprise, we got something that is fully functional,” Winther says, and retains perfect function from pH 2 to pH 12.
Emil Alexsov of Clemson University cautions that just because the uncharged protein was functional in a test tube assay doesn’t mean that it would work in a cell. However, the study does provide important insight on the effect of charge on solubility in water. In an uncharged protein, “solubility should decrease, but this was not the case,” he says.
Winther says that this study provides a blank slate for which to study charge interactions in proteins. “The next step is to introduce charges at different distances to each other or that are more or less buried in the structure,” he says, to define a charge’s sphere of influence, an important consideration for protein design.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter