Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Enzyme forms complexes with fractal geometries
Structures may result from an evolutionary accident
by Celia Arnaud, special to C&EN
April 24, 2024
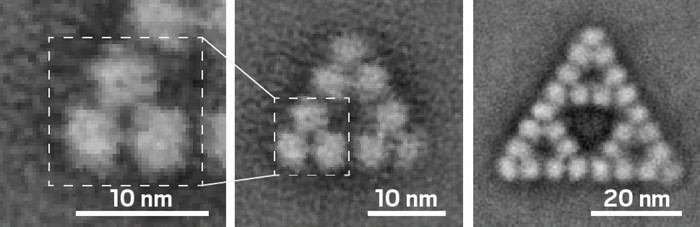

Researchers have identified an enzyme that can assemble into complexes with fractal geometries. Fractals—hierarchical patterns in which structural features at larger scales repeat at smaller scales—are well known at the macroscopic level but hadn’t previously been observed to spontaneously form from biological molecules at the molecular level in cells or in vitro.
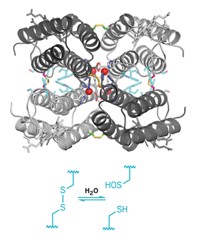
Now Georg K. A. Hochberg of the Max Planck Institute for Terrestrial Microbiology and the Philipps University of Marburg, Jan M. Schuller of the Philipps University of Marburg, and coworkers have shown that the enzyme citrate synthase from the cyanobacterium Synechococcus elongatus forms complexes in a fractal pattern called a Sierpiński triangle (Nature 2024, DOI: 10.1038/s41586-024-07287-2). Sierpiński triangles consist of small equilateral triangles nested within larger equilateral triangles.
The catalytic form of the S. elongatus citrate synthase is a triangular hexamer. These hexamers can assemble into Sierpiński triangles with 18 or 54 copies of the protein (3 or 9 of the hexamers). To form the fractals, the enzyme rotates in a direction opposite the way it rotates to bind its substrate during catalysis. The fractal “clamps the thing down in a way that makes catalysis difficult,” Hochberg says.
The enzyme forms these larger structures only at night, when the cyanobacterium’s pH is approximately neutral. “It’s possible that this thing is sort of a harmless accident, because it only makes this crazy structure at a time of day when you don’t need the enzyme anyway,” Hochberg says. The 18-mer forms at such low concentrations that Hochberg is convinced it happens in cells. He thinks the 54-mer is probably not formed physiologically.
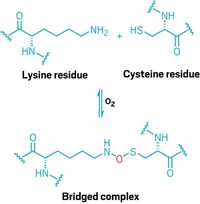
The researchers used ancestral protein reconstruction to investigate how the enzyme may have evolved the ability to form fractals. A glutamic acid and a histidine that are essential to the fractal-forming interface had been present in ancestral proteins that don’t form fractals. Replacing a glutamine with a leucine removed an interaction that had prevented fractals from forming. This change triggered them to assemble.
“That’s weird from an evolutionary perspective,” Hochberg says. “What it means is that all the positive contacts that glue this thing together were already there.”
“This is a fascinating example of how the vagaries of evolution can lead to the formation of architectures that would be challenging to realize via protein design because both interfacial contacts, steric clashes, and angular flexibility would have to be programmed in the hierarchy of noncovalent interactions that a single building block experiences as it assembles into a fractal,” François Baneyx, who engineers protein-containing materials at the University of Washington, writes in an email.
Eliminating the enzyme’s ability to form the fractals had no observed effect on the cells, Hochberg says. “These things are so shockingly easy to produce for evolution in a single mutational step that we should, in fact, expect that that sometimes happens by chance,” he says. If someone finds a similarly strange assembly in another organism, they might consider “whether this could just be a harmless accident,” Hochberg says.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter