Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
Aptamers Take on Proteomics
by CELIA M. HENRY, C&EN WASHINGTON
March 29, 2004
| A version of this story appeared in
Volume 82, Issue 13
Antibodies are usually considered the kings of affinity interactions. However, a type of nucleic acid molecule may turn out to be even better than antibodies, and these "aptamers" are finding use in all the applications that antibodies are known for. Some of these applications, especially genomics and proteomics, were the subject of a symposium at Pittcon.
Aptamers, sometimes called nucleic acid antibodies, are selected for their ability to bind other molecules, such as proteins, through a process of molecular evolution from a random pool of sequences. Aptamers were discovered independently by the team of Larry Gold and Craig Tuerk at the University of Colorado and that of Jack Szostak and Andrew Ellington at Harvard University.
Speaking at Pittcon, Gold, now the chairman and chief scientific officer at SomaLogic in Boulder, Colo., said that aptamers are more potent and more specific than antibodies. The large interface between the aptamer and its target protein partially explains the strong binding between the two molecules. The polyanionic character of the aptamer doesn't "wreck" the binding interface, Gold said.
SomaLogic is using aptamers to create protein arrays for proteomic applications, particularly the identification of surrogate biomarkers of disease. Such applications require limits of detection below the picomolar level, Gold said, because most protein variations correlated with disease occur in proteins expressed at low levels.
One of the ways SomaLogic can achieve such low detection levels is through the use of photoaptamers as capture agents. These molecules use the standard nucleotides based on adenine, cytosine, and guanine, but thymine is replaced by a nucleotide known as 5-bromodeoxyuracil. After the aptamer binds its target, ultraviolet light is used to generate a free radical, which attacks certain amino acids to form a covalent bond and cross-link with the protein. The cross-linking makes the binding permanent and increases the specificity of the interaction.
SomaLogic is using its aptamers in assays called antibody-linked oligonucleotide assays, or ALONA. In this sandwich-type assay, the aptamers bind to their target proteins, are cross-linked, and are then washed stringently. Detection is achieved with a fluorescently labeled antibody that binds to the cross-linked complex.
The aptamers also can be used in nonsandwich assays using a universal protein stain. This dye covalently attaches to proteins at the amino side group of lysine. The chemical attack allows SomaLogic to quantify how much protein is bound to the aptamer array without having to worry about a background signal because the dye doesn't bind to any free aptamer.
Ellington, now a professor of biochemistry at the University of Texas, Austin, would like to see aptamers become widely used, much in the way that array technologies developed by Patrick Brown at Stanford University caused complementary DNA microarrays to proliferate. Aptamers could similarly be spotted on arrays.
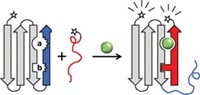
Weihong Tan, a chemistry professor at the University of Florida, Gainesville, turned to aptamers when the molecular beacons he was using for gene recognition studies turned out not to work for proteins. Molecular beacons are stem and loop nucleic acid structures with a fluorescent dye and quencher attached to the ends of the stem. When the molecular beacon interacts with its target, the beacon undergoes a conformational change that causes the quencher to move away from the dye, thus turning on the fluorescence. This method is known as fluorescence resonance energy transfer, or FRET.
"WE TESTED a couple of proteins with existing molecular beacons," Tan said. "We found that the reaction is very fast and the sensitivity is very good, but there's no specificity of recognition of a protein. Then we turned to what we call molecular beacon aptamers."
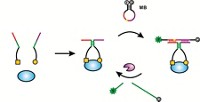
Tan combines the signaling transduction of molecular beacons with the specificity of aptamers. Depending on the type of interaction between the aptamer and its target, three different signaling methods are possible, Tan said. If there is a large conformational change associated with the binding, Tan can use the fluorophore and quencher or two-fluorophore approach to FRET. Using two fluorophores increases the signal and also allows for ratiometric measurements, such as the ones that are used with messenger RNA arrays.
If the aptamer-protein interaction does not involve a large conformational change, Tan uses fluorescence anisotropy (the difference in fluorescence intensity in the vertical and horizontal directions) to detect the interaction. This method requires that the aptamer be labeled with only one fluorophore. Tan takes advantage of the large size difference between the free aptamer and the aptamer bound to its protein target, which causes a large difference in the anisotropy measurements.
If feasible, FRET is preferable because it provides more sensitivity. But "anisotropy will always work," Tan said. He is using anisotropy to do intracellular imaging of proteins in living cells.
Tan is also looking at a new way to select aptamers. Rather than finding one aptamer at a time, he wants to find groups of aptamers that are selective for one type of cell (tumor cells, for example) over another (normal cells). In this approach, a pool of random DNA sequences is mixed with the cancer cells. The sequences that bind to the cancer cells are removed and then tested with normal cells. The aptamers that remain unbound in the second pool are specific for cancer cells over normal cells. Such an approach will allow Tan to find panels of protein markers that he hopes to use for diagnostics.
Ellington would like to use aptamers without the need for a label. He is collaborating with Scott Manalis at Massachusetts Institute of Technology to develop aptamer-based microcantilever biosensors. In these devices, interdigitated fingers from parallel cantilevers are used to make an interferometer.
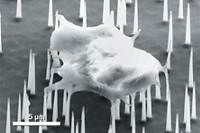
Despite evidence showing that aptamers are even more potent and selective than antibodies, scientists don't really understand what characteristics provide these benefits. Xiaohong Fang, a chemistry professor at the Institute of Chemistry at the Chinese Academy of Sciences, Beijing, is using atomic force microscopy (AFM) to better understand the interaction between the aptamer and its target.
Fang pointed out that although many papers claim that aptamers will rival or even replace antibodies in many applications, very little work has been done directly comparing the two. She presented data comparing the binding of three model proteins with aptamers and with antibodies. AFM measurements show that the binding force is higher with the aptamer, indicating a higher affinity. The researchers also are using AFM to study the binding properties of protein to aptamer and to explore the energy landscape of protein-aptamer complexes at the molecular level.
In addition, Fang described a method of using a label-free aptamer for protein detection, with the help of what she called a "DNA molecular light switch." The switch is a ruthenium complex that intercalates with DNA. Water causes its luminescence to be quenched. In the absence of the protein target, the complex is protected by intercalating with a DNA or RNA aptamer. When the protein binds to the aptamer, the luminescence decreases, most likely because the binding distorts the aptamer, releasing the ruthenium complex. This simple method avoids the need for designing an aptamer-labeling strategy or for coupling fluorophores to aptamers.
In addition to being used for selective protein detection, aptamers can also be used in other affinity applications. Robert T. Kennedy, a professor in the departments of chemistry and pharmacology at the University of Michigan, described his work developing stationary phases for affinity chromatography with aptamers.
Kennedy sees aptamers being used in proteomics and metabolomics as the first dimension in a multidimensional separation, in which the aptamers are selected to weakly bind to a class of proteins or metabolites. Other compounds would simply move through the column without being retained.
For affinity chromatography in which the column is being used to separate analytes, low-affinity aptamers are sufficient. "You don't want picomolar affinity," Kennedy said. "You want micromolar affinity. You don't want [the analyte] to get stuck there forever."
In contrast, affinity chromatography for purification purposes calls for higher affinity aptamers. "If you're really trying to preconcentrate something or purify something, then high affinity is good," Kennedy said. "You want high affinity and high capacity so you can inject large volumes of dilute solution, selectively retain your analyte, and then knock it off with some kind of elution mechanism. You want at least low nanomolar affinities to make that work well."
For chromatography applications, the aptamers don't need to be labeled. In capillary electrophoresis (CE) applications, the ease of labeling aptamers makes them attractive for analysis of complex mixtures. Kennedy sees the aptamer in that case as a targeted labeling technique.
His group found that CE with aptamers requires using the shortest column and lowest electric field possible to achieve the separation. They found that high electric fields speed up the separation but have the unfortunate side effect of pulling the aptamer-analyte complex apart.
HE SPECULATED on two reasons that might be happening. If a complex dissociates at low electric field, it still has a chance of recombining by diffusion. On the other hand, complexes that fall apart in a high electric field have very different velocities, so the aptamer and analyte never have a chance of recombining. The other possible cause of the complexes falling apart at high electric fields is the increased friction as the complex moves through the solvent at higher velocity. In the second scenario, the electrophoresis itself induces the complex to fall apart.
Kennedy would like to use aptamers to improve separations of complex biological mixtures, such as serum samples and brain dialysate samples. He described a chromatographic separation of adenosine-related compounds from a brain dialysate sample using an aptamer-based chromatographic column. Adenosine is a neurotransmitter involved in regulating sleep.
"A big part of our research is to use rapid separations as a monitoring application. We couple dialysis sampling probes in the brain to CE systems," he said. "We'd like to have aptamers for peptide hormones or peptide neurotransmitters and be able to detect them rapidly with this approach in an automated serial fashion."
So far, Kennedy has focused on aptamers that have already been discovered and are readily available. He realizes, though, that using aptamers for new applications will require raising them himself. "You can only go so far with taking aptamers that have been published ... and using them for your own [applications]," he said. "You can use them for proof-of-concept kinds of things, but if you really want to go for some proteomic separation or metabolomic application, you're going to have to [come up with] your own."
MORE ON THIS STORY







Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter