Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Synthesis
Electron-Starved Enzyme
Cytochrome c oxidase model mimics natural electron-limited conditions
by Celia Henry Arnaud
March 19, 2007
| A version of this story appeared in
Volume 85, Issue 12

A new model of the active site of a key enzyme in cellular respiration allows scientists to study the enzyme under conditions where the flow of electrons is the limiting step, as it is in the natural enzyme (Science 2007, 315, 1565).
Cytochrome c oxidase catalyzes the four-electron reduction of O2 to H2O during the final stage of respiration. This reduction must happen without releasing partially reduced oxygen species, which are toxic. How the enzyme accomplishes this is poorly understood, because scientists haven't been able to study the enzyme under electron-limited conditions that might be expected to lead to partial reductions.
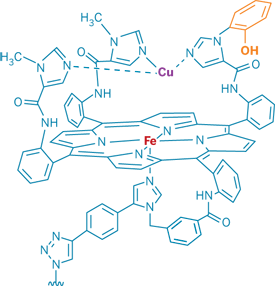
"The enzyme is always starved for electrons, something few people seem to recognize," says chemistry professor James P. Collman of Stanford University. Collman worked with graduate student Neal K. Devaraj, research associate Richard A. Decréau, and coworkers to build a biologically relevant model system that includes three redox sites: a myoglobin-like heme, copper suspended among three imidazoles about 5 Â above the heme, and a phenol group covalently attached to one of the copper-ligating imidazoles.
In earlier models, Collman and coworkers adsorbed the catalysts on graphite electrodes, so the electron transfer was very rapid. Now, in collaboration with Stanford chemistry professor Christopher E. D. Chidsey, they slow the electron transfer by fastening the catalyst covalently onto a gold electrode modified with a self-assembled monolayer.
The anchored model system answers some elusive questions about cytochrome c oxidase's mechanism, Collman says. "Biologists have usually studied cytochrome c oxidase by loading it with electrons and allowing it to turn over one time. It's really difficult for biologists to study cytochrome c oxidase under steady-state conditions and see what happens right at the active site," he says.
For example, biologists have suspected that all three of the natural enzyme's redox sites were necessary for preventing the release of partially reduced oxygen species, but it's been difficult to demonstrate. "By protecting in turn each of the redox-active substituents, we're able to show that to get the four-electron reduction that cytochrome c oxidase must do in order to prevent the formation of these toxic partially reduced oxygen species, all three substituents must be present at once," Collman explains.
Constructing the model has been a long-term effort, Collman says, culminating with the addition of the crucial phenol, which mimics a key tyrosine residue in the natural enzyme. Constructing the model requires a 32-step convergent synthesis.
The model doesn't mimic everything the natural enzyme does, Collman points out. With a fully oxidized enzyme, the first electron goes to a peripheral redox site outside of the active site. When a second electron is added, a conformational change causes both electrons to jump to the active site. In this way, the enzyme avoids forming partially reduced oxygen species. "We have not been able to imitate that," Collman says. "That's something we would like to do."


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter