Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Faithful Mimic of Proteins
Selective chemistry enables construction of modified proteins
by Celia Henry Arnaud
April 30, 2007
| A version of this story appeared in
Volume 85, Issue 18
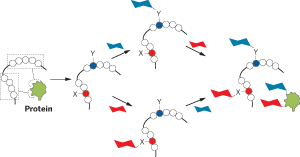
IN WORK that should help investigators unveil the structure-function relationships of proteins, researchers have chemically modified the bare scaffold of a natural protein such that the product functions like a different protein.
Many proteins undergo chemical changes after their amino acid chains, or bare scaffolds, are constructed in the ribosome. The process, called posttranslational modification (PTM), installs on the scaffold various groups, such as sugars and phosphates, without which the protein couldn't perform its unique function.
Because the same scaffold can be varied by PTMs in countless ways, PTMs increase protein diversity and could help explain biological diversity. Teasing apart the relationship between PTMs and specific protein function has been difficult so far, however, because researchers have not been able to chemically execute combinations of PTMs effectively.
Now, Benjamin G. Davis and coworkers at the University of Oxford report a chemical-tagging approach that allows them to attach multiple groups to a bare protein scaffold (Nature 2007, 446, 1105). The additions act like the natural modifications they are designed to mimic even though they are linked to the protein differently.
Davis and coworkers make modifications to specific locations on the scaffold by incorporating cysteine or by replacing methionine with nonnatural analogs containing azide or alkyne groups. The researchers then install sugars and other groups by taking advantage of selective chemistries involving the sulfide group in cysteine and the azide or alkyne groups on the methionine analogs. The reactions form the same products regardless of the order in which they are performed. And the chemistries are efficient enough that reactants are completely converted to the desired products.
Davis' team used the approach to mimic the human protein P-selectin-glycoprotein-ligand-1 (PSGL-1). This is a transmembrane protein found in white blood cells. Its target, P-selectin, is involved in the inflammatory response. Previous studies have shown that two PTMs are key to PSGL-1's function.
The researchers showed that they could install functional mimics of those PTMs on an unrelated protein scaffold and still get the desired function. For the scaffold, they used a protein with galactosidase activity called SSβG. By coupling this activity with the release of a blue dye, they could detect the scaffold's binding. They used structural information to figure out where on the SSβG scaffold to install the two modifications, a sugar and a sulfated side chain. The modified protein binds P-selectin in animal models of inflammation and malaria, the researchers report.
"By using our mimic, we could visualize inflammation at really early stages of diseases," Davis says. "The tagging strategy is really quite conceptually simple; the exciting thing is that we can make a synthetic mimic that turned out to be a much better way of visualizing this than corresponding antibody methods."
Carolyn R. Bertozzi, a chemistry professor at the University of California, Berkeley, calls the work "an elegant illustration of how chemical synthesis can produce biological probes that would be impossible to derive by conventional biochemical or genetic methods."


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter