Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Analytical Chemistry
DNA Repair Mechanism Probed
Single-molecule methods, crystallography reveal surprises about strand-swapping process
by Celia Henry Arnaud
June 16, 2008
| A version of this story appeared in
Volume 86, Issue 24

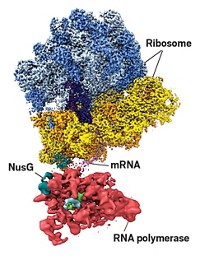
MANY BIOLOGICAL SITUATIONS call for swapping segments of DNA between strands. This process, known as homologous recombination, is vital for repairing broken DNA and generating genetic diversity. Although biologists have studied this process for many years, aspects of it have remained cloaked. Two new studies uncover different portions of the reaction.
In homologous recombination in bacterial systems, a piece of single-stranded DNA (ssDNA) forms a filament coated with an enzyme called RecA, which catalyzes the insertion of the ssDNA into a DNA duplex and the expulsion of one of the original DNA strands. Adenosine triphosphate (ATP, the main energy-transfer molecule in the cell) is hydrolyzed in the process.
Cees Dekker and coworkers at Delft University of Technology, in the Netherlands, used single-molecule methods to watch homologous recombination in real time (Mol. Cell 2008, 30, 530). They find that no matter how long the piece of DNA to be inserted, the synapse—the portion of the complex where RecA binds all three DNA strands—is always approximately 80 nucleotides long. The synapse "could have been zero, or it could have been kilobases," Dekker says. "We find a small but finite length."
Dekker and coworkers used a manipulation tool, magnetic tweezers, to control the topology of the DNA on either side of the synapse. In this technique, one end of the double-stranded DNA (dsDNA) is attached to the bottom of a flow cell. The other end is tethered to a magnetic bead. By manipulating a pair of magnets held above the flow cell, the scientists then can twist and stretch the immobilized molecule.
Precise measurement of the location of the magnetic bead reveals the end-to-end distance of the DNA molecule, which changes as the RecA-coated ssDNA filament interacts with the tethered DNA duplex. As the ssDNA incorporates itself into the DNA duplex, the end-to-end length of the complex increases linearly with time, but the increase levels off at 80 nucleotides. This length change is the same whether Dekker's group used an invading strand that is 359 nucleotides long or 1,055 nucleotides long.
Dekker says his data suggest that the synapse forms a structure that he calls a "D wrap." The name derives from the related structure, called a D loop, formed when a DNA duplex is pulled apart and one strand of the resulting bubble is paired with complementary DNA. The tight topological control of Dekker's molecular tweezers protocol won't let the typical D loop form. "We have already twisted the DNA in a certain way," he says. When the duplex is pulled apart and the invading strand binds, the expelled strand wraps itself around the newly formed duplex.
Dekker suspects that the D wrap will be applicable in cellular settings as well. Chromatin, the DNA-protein complex that makes up chromosomes, is often similarly topologically constrained upstream and downstream of the recombination location, he says. People may not have seen the D wrap structure before, he says, because many experiments have been performed with freely rotating DNA oligomers.
Taekjip Ha, a biophysicist at the University of Illinois, Urbana-Champaign, points out that this is not the first time strand exchange has been observed at the single-molecule level. But "it is the cleanest experiment," he says. He finds the constant length of the synapse interesting and important but notes that some people may think that length is too long because strand exchange reactions have been performed in the past with shorter oligonucleotides.
HOMOLOGOUS recombination is a "classical molecular biology textbook reaction," says Antoine van Oijen, a professor at Harvard Medical School who develops single-molecule methods and uses them to study biophysical processes. "Through decades of great structural, genetic, and biochemical work, we know a lot about the mechanism, but certain details have remained hazy. I think it is a great example of how the single-molecule field is growing up and how these techniques are able to address those key aspects of biochemical pathways that have been difficult to address using bulk-phase methods," van Oijen says.
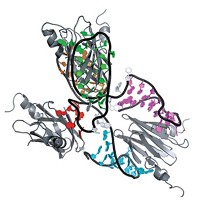
Recent crystallographic work has cleared up other details of the mechanism of homologous recombination, particularly how RecA interacts with DNA. Structural biologist Nikola P. Pavletich and coworkers at Memorial Sloan-Kettering Cancer Center, in New York City, report crystal structures of RecA with ssDNA and dsDNA (Nature 2008, 453, 489). Pavletich's team obtained filament structures that can exchange strands only with DNA bound to fused proteins made of five or six RecA units.
Pavelich's crystallographic report follows a study in which Ha's group used single-molecule methods to show that RecA nucleation on ssDNA requires five protein monomers (Cell 2006, 126, 515). Ha suggests that the current work "could be considered an example where a finding in single-molecule studies motivated a new research direction in a more traditional biological research enterprise."
Pavletich's structures suggest that the interactions of ssDNA and ATP with RecA are allosterically coupled. Binding of the ssDNA and ATP shifts the RecA conformation in a way that creates a new interface between RecA subunits and activates the filament for strand exchange.
Obtaining these crystallographic structures was "a tremendous achievement," Ha says. "People have tried to obtain the cocrystal structure for more than 15 years."


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter