Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Synthesis
Build Your Own Enzyme
Biochemistry: Scientists create the first intermolecular Diels-Alderase
by Bethany Halford
July 19, 2010
| A version of this story appeared in
Volume 88, Issue 29
When it comes to catalysis, chemists have nothing on nature’s enzymes, which get reactions to go smoothly and stereoselectively under the mildest of conditions. Now, taking the “if you can’t beat them, join them” approach, scientists have designed and built enzymes from scratch that can catalyze an intermolecular Diels-Alder reaction—a transformation no naturally occurring enzyme is known to do (Science 2010, 329, 309).
In the past, engineered enzymes have only been able to perform simple bond-breaking reactions. The new work, spearheaded by David Baker of the University of Washington, Seattle, demonstrates that de novo-designed enzymes can do more complex chemistry; in this case, forming two carbon-carbon bonds.
And because intermolecular Diels-Alder reactions are used to make myriad chemicals, including pharmaceuticals, materials, and fine chemicals, enzymes that can make those reactions go more efficiently or under greener conditions than currently possible could find widespread use, says Justin B. Siegel, one the report’s primary authors.

“It opens a whole new dimension for catalyst design for multicomponent condensations using protein engineering,” says John C. Vederas, a chemistry professor at the University of Alberta, Edmonton. “It is especially exciting because the system initially binds and holds the two reaction partners in a desired three-dimensional orientation with respect to one another, something that is difficult to achieve predictably in standard solution chemistry.”
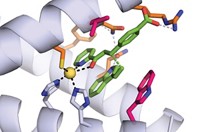
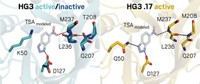
Baker’s team chose to build an enzyme for catalyzing the Diels-Alder reaction of the diene 4-carboxybenzyl trans-1,3-butadiene-1-carbamate and the dienophile N,N-dimethylacrylamide. Using Rosetta computational design methodology, the researchers prepared an in silico model of the shape that would be needed to accommodate the transition state for this reaction. They then added amino acids that would hold the reactants in place—a hydrogen-bond acceptor to interact with NH on the diene’s carbamate and a hydrogen-bond donor to interact with the dienophile’s carbonyl.
“Once we have the shape and chemistry, then we have to fill in the rest of this binding pocket,” Siegel explains. Calculations indicated that 1019 theoretical active sites matched the stipulations the group had set out. They then used the RosettaMatch program to screen these possibilities against known protein scaffolds and determined that 106 would correspond to a stable scaffold.
Further computational modeling and scientific intuition narrowed the number of possible enzymes to 84, all of which the researchers expressed and purified. Of those, 50 turned out to be soluble and two had Diels-Alderase activity. Fine-tuning of the amino acids in the active sites of those two enzymes boosted their activity so that they performed better than catalytic antibodies designed to perform the Diels-Alder reaction, although Baker cautions that a designed enzyme isn’t nearly as catalytically active as a native enzyme. Further studies on one of the de novo Diels-Alderases showed both stereoselectivity and substrate specificity.
“There is endless fascination with the Diels-Alder reaction owing to its topological elegance and utility in organic synthesis. Its apparent lack of discovery by nature adds additional intrigue,” Yale University’s William L. Jorgensen comments. The Baker team’s success, he says, “represents a great advance in the latter regard and for rational enzyme design.”



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter