Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Unnatural Bases Help Scientists Mark DNA Lesions
DNA Analysis: Hydrophobic base pair works with PCR amplification and nanopore sequencing to tag and identify DNA damage
by Celia Henry Arnaud
November 12, 2015
| A version of this story appeared in
Volume 93, Issue 45

DNA mutations can cause many diseases such as cancer. But by the time scientists identify DNA mutations, the original chemical damage that caused them is lost.
“What we’d really like to do is sequence for the chemical modifications, the origins of the mutations,” says Cynthia J. Burrows, a chemistry professor at the University of Utah.
Unfortunately, damaged nucleotides block the enzymes needed to amplify DNA for sequencing. Burrows and coworkers have overcome this barrier with a method that amplifies and detects DNA lesions. To do that, they mark DNA damage with an unnatural base pair that is compatible with the polymerase chain reaction, or PCR (Nat. Commun. 2015, DOI: 10.1038/ncomms9807).
First they use enzymes from the DNA base excision repair pathway to identify and clip out the damage, leaving a gap in the DNA. On the basis of which enzymes they end up needing, the team can determine the kind of damage that was present.
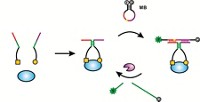
Next, Burrows and coworkers fill the gap with hydrophobic unnatural bases developed by Floyd Romesberg at Scripps Research Institute, La Jolla, Calif. “We just give it one nucleotide in relatively high concentration and a polymerase enzyme,” Burrows says. Such conditions force the polymerase to incorporate the unnatural nucleotide, even though it’s a mismatch with the base sitting across from the gap.
Then the researchers amplify the mismatched duplex using PCR. By including a second unnatural nucleotide that is fully complementary to first, they are able to generate many copies of DNA with the unnatural base pair.
Biochemists previously showed that polymerases could incorporate hydrophobic nucleotides across from gaps in a DNA sequence, says Eric T. Kool, a chemistry professor at Stanford University who studies unnatural nucleotides. “However, the new work takes an important practical step ahead by employing a full base pair that can be amplified by PCR.”
Finally, the Utah team takes those DNA copies and uses protein nanopore sequencing to identify the location of the unnatural base, which tells them where the lesion was originally. To make the unnatural bases easy to detect, they label them with a crown ether that modulates the current measured through the nanopore.
“The diversity of lesion structures and their exceedingly low abundance in the genome makes it seem like an impossible problem” to spot and characterize them, says Shana J. Sturla of ETH Zurich, who previously developed a method for marking a particular type of lesion. “By developing a clever strategy that brings together different enzymological and chemical tricks, whereby the lesion can be found, marked, amplified, and detected, Burrows and coworkers start to make the impossible seem possible.”


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter