Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Improved route to single-base genome editing
Modified CRISPR-Cas9 technique corrects disease point mutations by efficiently changing single bases in DNA
by Stu Borman
April 21, 2016
| A version of this story appeared in
Volume 94, Issue 17

In a new approach to genome editing called “base editing,” researchers have engineered the CRISPR-Cas9 enzyme to modify individual DNA bases more efficiently and more accurately than with existing gene-editing methods.
Many genetic diseases arise from single-base mutations, or point mutations. David R. Liu, postdoc Alexis C. Komor, and coworkers at Harvard University developed the new base-editing process and showed that it can correct point mutations in living cells, suggesting the method could lead to therapies that correct genetic errors that cause human diseases (Nature 2016, DOI: 10.1038/nature17946).
Researchers currently use zinc finger nucleases, transcription-activator-like effector nucleases, and CRISPR-Cas9 for genome editing. All three tools initiate genome editing by cutting both strands of DNA. Cellular machinery reduces the efficiency of these editors’ ability to correct point mutations by trying to repair such double-strand breaks with so-called indels, random DNA insertions or deletions.
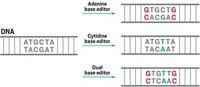
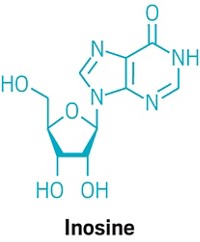
By avoiding double-strand breaks, the new approach causes far fewer indels and corrects point mutations more efficiently. Liu and coworkers began by using a Cas9 mutant (dCas9) that doesn’t cut DNA but retains native Cas9’s ability to bind to target DNA sequences. In this case, the team wanted to target C:G base pairs and change them to T:A. They made their first base editor by fusing the mutant to cytidine deaminase, which converts cytosine (C) to the RNA base uracil (U), thus changing targeted C:G base pairs to U:G. Cellular DNA repair and replication machinery can correct such U:G mismatches to U:A or T:A.
But cells’ DNA repair machinery can also change U:G back to C:G, so the researchers made a second base editor by appending to their first base editor a small protein called uracil glycosylase inhibitor, which prevents that switchback. In human cells, the second base editor increased the efficiency of base editing several-fold, compared with the first. Liu and coworkers then restored one of the DNA-cutting active sites in dCas9, creating a third base editor that further increases efficiency by favoring cellular repair of U:G to U:A or T:A over the switch back to C:G.
In tests in cell culture, Liu and coworkers observed that conventional CRISPR-Cas9 corrected 14 point mutations about 1% of the time, while causing about 5% indels. Their second base editor corrected about 10% of the same mutations and caused almost no indels, whereas the third was about 30% efficient and caused about 1% indels. Both the second and third base editors are useful, Liu notes, depending on whether mutation efficiency or avoiding indels is the more important goal of a particular experiment.
The researchers demonstrated the power of the technique by using it to reverse point mutations associated with Alzheimer’s disease and cancer in mouse and human cell lines, with minimal indel formation.
Emmanuelle Charpentier of the Max Planck Institute for Infection Biology, who helped discover the CRISPR-Cas9 technique, comments that “the study shows once again the power and versatility of the CRISPR-Cas9 mechanism.” The work “predicts future exciting applications in tailored genome and epigenome base editing.”
This article has been translated into Spanish by Divulgame.org and can be found here.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter