Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Designed peptides with constrained structures are good drug prospects
Computational modeling technique creates peptides with fixed, unprecedented shapes
by Stu Borman
September 19, 2016
| A version of this story appeared in
Volume 94, Issue 37
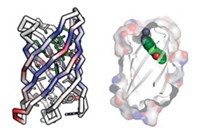
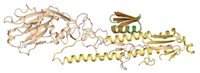
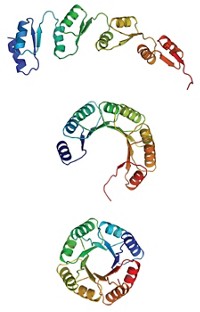
A newly developed computational approach makes it possible to design peptides with defined three-dimensional structures that could be useful for drug discovery (Nature 2016, DOI: 10.1038/nature19791). Peptides are often too floppy and undisciplined to interact with molecular targets in a consistent way. But the designed peptides’ disulfide bonds and terminal cyclization constrain them into defined shapes that enhance their potential for bioactivity. Protein design generally relies on prior knowledge of natural protein structures, but first-principles simulation techniques reported in the new study by David Baker of the University of Washington and coworkers optimize the creation of nonnatural peptides without reference to known structures. They used the system to design 16 peptides with unprecedented shapes, in some cases made from both natural and nonnatural amino acids. They created the designed structures by peptide synthesis or by expression in engineered bacteria. The designed peptides are resistant to thermal and chemical denaturation, and X-ray and NMR structures of 12 of them showed them to be nearly identical to their computational design models. “We are now working on developing peptides that target pharmacologically relevant proteins,” says Baker group postdoc Gaurav Bhardwaj.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter