Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Insect sodium channels visualized with cryo-electron microscopy
Structures of voltage-gated channel with toxins suggest strategies for drug development
by Bethany Halford
July 26, 2018
| A version of this story appeared in
Volume 96, Issue 31

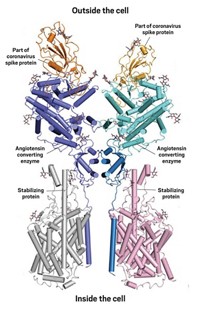
Scientists now have a better picture of how voltage-gated sodium (Nav) channels interact with certain animal toxins, thanks to structures solved with cryo-electron microscopy. These channels are important targets for insecticides as well as for drugs that affect cardiac function and pain.
Much of the structural data scientists previously had on Nav channels came from crystallographic studies on channels derived from bacterial proteins. These channels, which are composed of four identical subunits, are molecularly distinct from the Nav channels of higher organisms, which are a single chain that forms four regions, explains Nieng Yan, a Princeton University molecular biologist who led the new study.
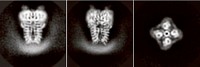
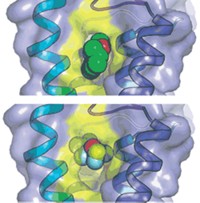
Yan’s group, along with researchers in Glenn F. King’s lab at the University of Queensland and in Qiang Zhou’s lab at Tsinghua University, used cryo-EM to create a picture of the American cockroach’s Nav channel, capturing details down to 2.8 Å resolution (Science 2018, DOI: 10.1126/science.aau2596). They solved the structure in the presence of a toxin derived from the desert bush spider—a peptide that keeps the channel open. They also determined the structure of the peptide-channel complex with the small molecule toxins tetrodotoxin (at 2.6 Å resolution), which is found in pufferfish and other critters, and saxitoxin (at 3.2 Å resolution), which comes from marine microorganisms and is responsible for certain types of shellfish poisoning.
Prior work has established that the small-molecule toxins block Nav channels, and the researchers’ structures show these molecules sitting snugly in the channel’s pore, which they expected, Yan says. What surprised them was the complex binding of the spider peptide toxin, which was previously thought to coordinate just one remote region of the channel but instead makes key contacts with the extracellular dome region of the channel that’s above the pore.
William A. Catterall, an expert in electrical signaling between cells at the University of Washington, says the work is “a tremendous step toward understanding the molecular and chemical basis for the powerful effects of animal neurotoxins on our nervous system that are caused by modulating voltage-gated sodium channels.”
“Because Nav channels are evolutionarily conserved from insects to humans, the findings here have enormous implications for development of pore blockers and gating modifiers specific to human Nav channels in order to treat chronic pain, epilepsy, cardiac arrhythmia, and other conditions more effectively,” adds George Wisedchaisri, who studies the pharmacology of membrane proteins at the University of Washington.
Yan says this report is just a starting point. “We are really intrigued by these channels and our long-term goal is to elucidate their whole working cycle,” characterizing the conformational changes that take place as the channel moves from one state to another. “It’s just like a miniature machine,” she says.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter