Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Neuroscience
Vast variability found in Alzheimer’s gene
Researchers find thousands of variants in amyloid precursor in diseased brains
by Megha Satyanarayana
November 28, 2018
| A version of this story appeared in
Volume 96, Issue 48

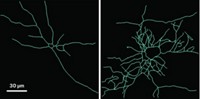
Neuroscientists have long wondered how a supposedly static genome could give rise to the brain’s incredible diversity in health and disease. Now researchers have found that a key gene implicated in the pathology of Alzheimer’s disease is highly variable. The findings could help explain outcomes as diverse as how we store memory, and in the case of Alzheimer’s disease, how we lose it (Nature 2018, DOI: 10.1038/s41586-018-0718-6).
Jerold Chun, a neuroscientist at Sanford Burnham Prebys Medical Discovery Institute and the University of California, San Diego, looked for variations in APP, the gene coding for the amyloid precursor protein. The buildup of amyloid-β, one cleavage product of the amyloid precursor protein, is thought to lead to neurotoxic plaques believed to cause Alzheimer’s disease.
Researchers knew APP had some variants, but the full breadth of variation found in the recent study was surprising, says Chun. The researchers studied APP messenger RNA and DNA sequences from brain tissue from people who have Alzheimer’s with no family history of the disease, and from people without Alzheimer’s. They found thousands of novel splicing variants. In people with Alzheimer’s disease, APP variants were more numerous and diverse.
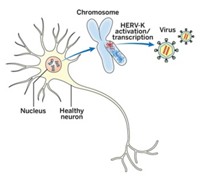
Chun and his colleagues also explored the mechanism behind APP variation. Some variants they found lack introns. This suggested to the group that neurons may use reverse transcriptase to make new versions of the gene. This enzyme is similar to the one HIV uses to copy itself in the human body, and that is targeted by HIV drug azidothymidine (AZT). When the researchers dosed cells with AZT, it seemed to reduce the production of APP variants. Chun wonders if people who are HIV-positive and on antiretroviral therapy might be protected from the variability in APP sequences and, consequently, from Alzheimer’s disease.
Chun speculates that APP’s variability may help explain why Alzheimer’s therapies have had limited success in clinical trials. Alternate toxic variants may take the place of the ones targeted by drugs. Chun says the ability to create these variations, if found in other genes in the brain, may help explain the organ’s diverse cell population and capabilities. This kind of genetic flexibility “appears to be a way you can change the very blueprint of a given neuron in response to activity,” Chun says.
The findings are intriguing, says Elizabeth Fisher, a neurogeneticist at University College London who studies Alzheimer’s disease in people with Down syndrome. Should they be reproduced by other labs, it would be a paradigm shift in how researchers understand the pathology of Alzheimer’s disease.
“It opens a cupboard of questions,” she says. “It potentially gives us a whole new set of cellular phenomena to investigate.”
But, above all, Fisher says, this research points to a dire need for neuroscientists to collect more frozen brain samples. The lack of starting material makes it challenging to repeat this and other studies, and even to do them in the first place, she says.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter