Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
RNA
Click chemistry–based RNA sequencing captures bursts of transcription
A new technique allows scientists to track RNA expression in single cells at the moment genes are transcribed from DNA to RNA.
by Max Barnhart
June 10, 2024

There are only about 20,000 different genes in the human genome, yet our bodies are filled with an incredible diversity of cell types performing innumerable functions. The key to that diversity in form and function relies on differences in how those genes are expressed. Now, scientists are one step closer to understanding how gene expression is regulated. MIT researchers have developed a new technique using click chemistry to capture which genes are in the process of being transcribed in single cells, which they say could be the key to better understanding and treating some genetic diseases (Nature 2024, DOI: 10.1038/s41586-024-07517-7).
RNA transcription isn’t a fully continuous process; instead it often occurs in short bursts where, at one moment, a gene may be highly expressed, but at another time, that gene may not be expressed at all. The timing of these transcriptional bursts is tightly coordinated with the activity of regulatory regions of the genome called enhancers. Phil Sharp, a cell biologist at MIT and author on the paper, says that this new methodology allows researchers to “in a very fine temporal manner, measure the activation of transcription” to see which genes are undergoing a transcriptional burst and how that relates to enhancer regions in a single cell.
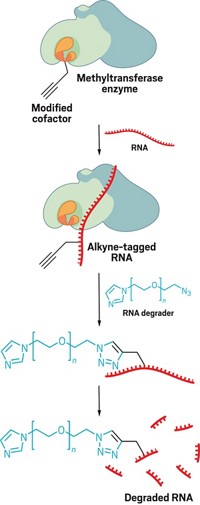
To carry out this method, the researchers feed cells modified nucleotides that contain a special alkyne group. During a burst of transcription, those modified nucleotides get incorporated into the RNA. Then the researchers isolate the nuclei from individual cells and use copper-catalyzed azide-alkyne cycloaddition click chemistry to attach a DNA barcode to the alkyne groups on the modified nucleotides. Each barcode is unique to a nucleus from a single cell, giving the researchers the ability to track exactly which genes are undergoing a burst of transcription in a specific cell.
Jay Mahat, a molecular biologist at MIT and lead author of the study, says that the click chemistry involved “lives up to its name. It’s very efficient in multiple conditions and easy to work with.” Pablo Mendieta, a functional genomicist at the University of Georgia who wasn’t involved in the research, agrees, calling the use of click chemistry in single cell sequencing “really innovative” and adding that “there’s a really large utility here to measure [active] transcription at a low molecule count, which is really hard.”
Sharp says that gene regulation underlying some genetic diseases, including sickle cell anemia, is “driven by enhancers” and that this methodology can help to better understand how those enhancers play a role in these diseases. Mahat adds, however, that researchers need to develop a more high-throughput version before the full potential of this technique can be realized. “Right now, we can process thousands of cells. Ideally you want tens of thousands of cells,” he says.
Mendieta also believes more testing is necessary. “We don’t know how well this will apply to diverse cell types,” he says, noting that only mouse embryonic stem cells were used in the paper.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter