Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Structural Biology
Delving into bacterial gene-expression machinery
Transcription factors connect and stabilize bacterial transcription and translation machinery
by Celia Henry Arnaud
August 20, 2020
| A version of this story appeared in
Volume 98, Issue 32
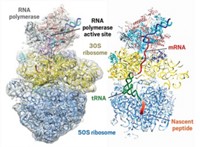
Three new studies provide structural clues about how ribosomes and RNA polymerase are coupled to one another during bacterial gene expression.
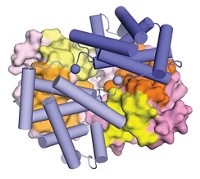
One team led by Albert Weixlbaumer of the Institute of Genetics and Molecular and Cellular Biology and another team led by Richard H. Ebright of Rutgers University used cryo-electron microscopy to image complexes called expressomes assembled from purified components produced by Escherichia coli (Science 2020, DOI: 10.1126/science.abb5036 and DOI: 10.1126/science.abb5317).
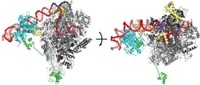
Both teams found that when the mRNA spacer between RNA polymerase and the ribosome active center is short, the components form a complex that matches a previously reported one. But the current studies find that complex likely represents a stalled state in which the RNA polymerase and ribosome have collided.
With longer mRNA spacers, the resulting complex is functionally active. In this active state, a transcription factor called NusG forms a bridge between the ribosome and the RNA polymerase. In addition, Ebright’s team solved structures in which a second transcription factor, NusA, further stabilizes the complex.
In a third study, Julia Mahamid of the European Molecular Biology Laboratory, Juri Rappsilber of the Technical University of Berlin, and coworkers used cryo-electron tomography and cross-linking mass spectrometry to determine the architecture of active expressomes inside Mycoplasma pneumoniae cells (Science 2020, DOI: 10.1126/science.abb3758). The researchers found both NusG and NusA as part of the expressome in Mycoplasma, but only NusA appears to form a contact between the ribosome and RNA polymerase. The positions and orientations of the transcription factors were different from those found in vitro with E. coli components.
The studies “provide an important framework for future studies,” says Andrei Korostelev, an expert on translation regulation at the University of Massachusetts Medical School. Future work “will uncover how the coupling is regulated by different RNA polymerase and ribosome states and by transcription and translation factors,” he says.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter