Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Knockout Rats The Easy Way
Zinc finger nucleases create genetic deletions in mammals with high precision
by Aaron A. Rowe
July 27, 2009
| A version of this story appeared in
Volume 87, Issue 30
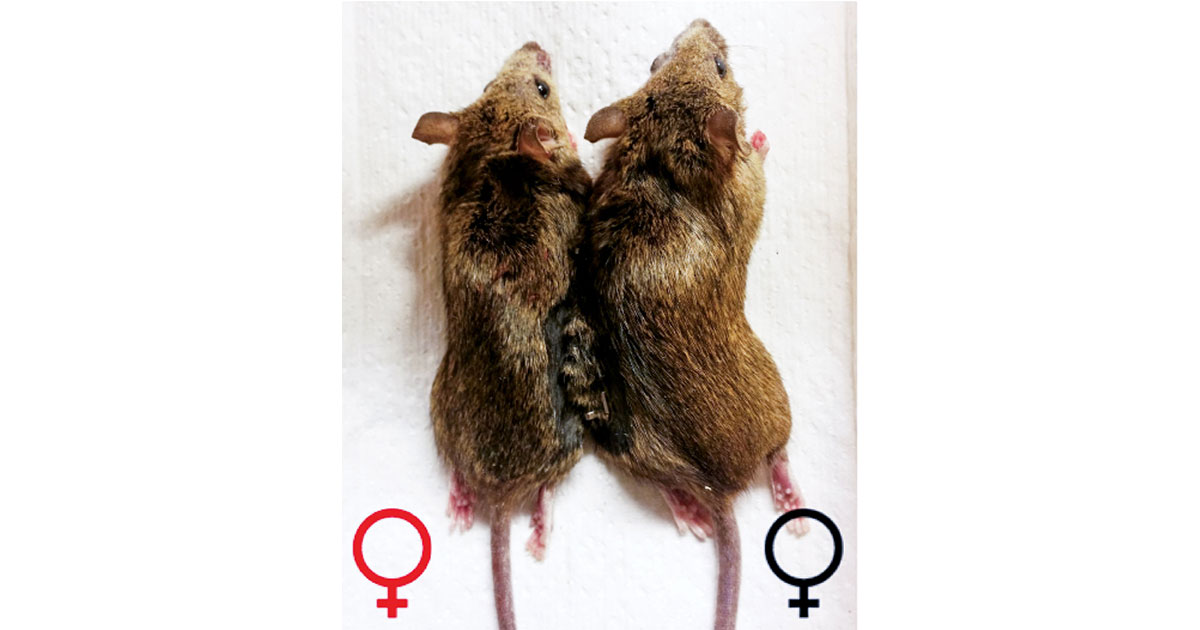
Groups of genetically engineered fluorescent rats have given birth to offspring that lack their surreal green glow. The pups’ appearances were altered in the embryonic stage of development by an engineered zinc finger protein that knocked out the gene that illuminated their parents, according to a new paper in Science (2009, 325, 433).
The new knockout strategy “is a more efficient method that allows the specific targeting of a gene,” says Michael N. Gould, the professor at the University of Wisconsin, Madison, who made the first knockout rats several years ago using a different technique. “In the future, this approach should also allow the ‘knock in’ of mutations within a gene.”
Roland Buelow of Open Monoclonal Technology and Howard J. Jacob from the Medical College of Wisconsin led the effort to create the new knockout rat pups.
Jacob hopes to use this new knockout strategy to build better animal models of disease. Knockout mice have become widespread tools for medical research, but rats are more similar to humans in some respects. Until now, creating rats that lack a particular gene has been challenging.
Buelow aims to use the knockout strategy to make transgenic rats that produce fully human antibodies. Since the paper was submitted, he has created rats that lack all three genes that are critical for the production of rat antibodies. By the end of this year, he hopes to have raised animals that bear human genes in their place.

The overall strategy relies on zinc finger proteins that can recognize and bind to exact DNA sequences. Sangamo BioSciences has built up an arsenal of tricks for engineering these natural proteins so that they bind to a target of choice. By attaching a FokI DNA-snipping domain to each zinc finger, the company can make nucleases that cause double-strand breaks in genes with pinpoint accuracy. When repair enzymes kick in, they sometimes shred a few bases from the ends of each damaged strand, which leads to mutations that render the genes inactive.
The biotech company has touted a wide range of applications for its flagship technology. It has begun a Phase I study of an HIV treatment and a Phase II trial of a remedy for diabetic neuropathy. And it recently clipped an unwanted gene from maize and replaced it with a code for herbicide tolerance.
Academics have complained that Sangamo is too protective of its intellectual property, but recently the company entered into a pact with Sigma-Aldrich to provide custom nucleases to researchers. Jacob received mRNAs that encode zinc finger nucleases from those companies and injected the proprietary genetic material into thousands of embryos. Within six months, the first set of engineered rats was born.
According to Buelow, the zinc finger nuclease technique could be used to make all sorts of knockout animals.



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter