Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biological Chemistry
Activation Of Signaling Protein Is Visualized
Structural Analysis: Study takes snapshots inside a working protein
by Stu Borman
November 19, 2012
| A version of this story appeared in
Volume 90, Issue 47

Researchers have devised a way to watch a light-induced signaling protein as it functions, revealing new details about the way it is activated to do its job. The technique could make it possible to better understand step-by-step mechanisms of other signaling proteins, which control processes such as the movement of leaves toward light and vision in higher animals, and possibly the mechanisms of enzymes as well.
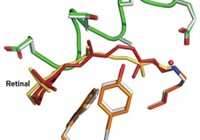
In the study, the researchers used time-resolved Laue crystallography to take structural snapshots of bacterial photoactive yellow protein (PYP) in action. When PYP is activated by potentially harmful light frequencies, it provides signals that induce the bacteria to swim away. Using the Laue method, which harnesses polychromatic X-ray pulses to rapidly obtain structural information from protein crystals, the researchers captured PYP’s activation with near-atomic spatial resolution and 150-picosecond time resolution.
Ultrafast biophysical chemistry specialist Philip A. Anfinrud of the National Institute of Diabetes & Digestive & Kidney Diseases, in Bethesda, Md., and coworkers carried out the study using a highly customized laser system on a synchrotron beamline at the Advanced Photon Source in Argonne, Ill. (Proc. Natl. Acad. Sci. USA, DOI: 10.1073/pnas.1210938109).
They visualized PYP activation at time delays ranging from 100 picoseconds to 1 second (a 10-order-of-magnitude range) after an initiating laser pulse. The technique’s 150-picosecond time window made it possible for them to view previously unobservable structural changes in a p-coumaric acid unit that undergoes trans-to-cis isomerization when the protein complex is activated. The study also visualized effects of hydrogen bonding, strain, and water in the activation process.
The study’s picosecond time resolution and the p-coumaric acid changes it revealed are important developments, says Laue crystallography expert John Helliwell of the University of Manchester, in England. But some of the findings on structural movements were close to the limits of observability, he notes, adding that he hopes scientists can extend those limits.
“One of the dreams of chemists, which I share, is to make molecular movies of structural motions,” comments Robin M. Hochstrasser, a structural dynamics specialist at the University of Pennsylvania. “Instead of taking spectra and making indirect interpretations,” Anfinrud and coworkers “directly measure scattering from atoms to watch a photochemical process evolve as a function of time. The work makes a beautiful connection between chemistry and biology.”
PYP is reversible—it resets itself after each signaling event and can therefore be probed repeatedly. “The next frontier,” Anfinrud says, “is to see if we can extend our methodology to systems we get only one shot at”—such as watching in real time and with near-atomic resolution a light-induced enzyme as it functions.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter