Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Proteomics
Unusual hydrogen bond plays a bigger role in proteins than chemists realized
Interaction affects 94% of all proteins, study shows
by Celia Henry Arnaud
October 19, 2016
| A version of this story appeared in
Volume 94, Issue 42
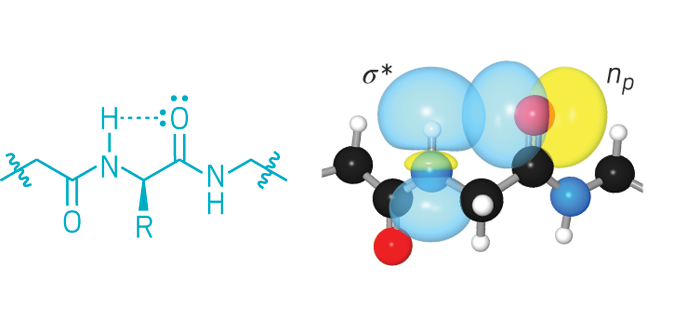
Since the 1960s, chemists have known about an unusual type of hydrogen bond: one that can form between the amide hydrogen and the carbonyl oxygen in the same amino acid. But it’s been hard to show that this interaction, called the C5 hydrogen bond, matters beyond amino acids, in proteins.
Now, biochemistry professor Ronald T. Raines and graduate student Robert W. Newberry of the University of Wisconsin, Madison, report experimental and computational evidence that this hydrogen bond indeed helps stabilize proteins—significantly in some cases (Nat. Chem. Biol. 2016, DOI: 10.1038/nchembio.2206).
“It has been problematic to quantify the energetic consequence of this H-bond, particularly in systems larger than a dipeptide where numerous other factors come into play,” says Steve Scheiner, a chemistry professor at Utah State University who studies hydrogen bonding. The new work “takes a major step toward unambiguously verifying the presence of this H-bond, as well as offering some quantitative measures of its influence.”
Raines and Newberry used spectroscopic measurements and quantum mechanical calculations of model peptides called tryptophan zippers, which are β hairpins that are 12 amino acids long, to show that the C5 interaction behaves like a hydrogen bond. They then used bioinformatics to analyze a database of protein structures, revealing that the interaction is widespread, affecting ~5% of amino acids and ~94% of all proteins.
The C5 H-bond is strongest in flat structures, which limits where it can occur. “It has to be in β sheets,” Raines says. “Otherwise, the oxygen and hydrogen are too far away from each other.” Raines speculates that this interaction could be important for stabilizing amyloid-forming proteins rich in β sheets, such as those involved in Alzheimer’s disease.
Incorporating this stabilization energy into models of protein folding could help improve predictions of protein structures, Raines says. “We’re very good at predicting the structures of things that are similar to what’s in the database,” he says. Predicting structures from scratch is much harder. “To truly solve the protein folding problem, we need to be much better in our accounting. We need to understand the fundamental forces that lead to a stable structure.”



Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter