Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Big Data
Big data technique predicts 12% of unknown protein structures
Milestone in modeling yields 100 new protein folds
by Sarah Everts
January 19, 2017
| A version of this story appeared in
Volume 95, Issue 4

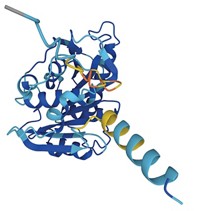
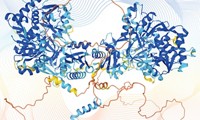
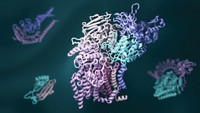
In a milestone achievement for computer-based prediction of protein structure, researchers have reported the three-dimensional conformations of 12% of the protein families that still have unknown structure. The cohort of 600 newly determined protein family structures includes 100 protein folds not found in the Protein Data Bank and 200 membrane proteins (Science 2017, DOI: 10.1126/science.aah4043).
A team led by David Baker of the University of Washington modeled the structures of these proteins using its Rosetta algorithm on a distributed network of volunteer computers. Baker’s team initially deposited some of the predicted structures it found into public databases. In the months afterward, structural biologists serendipitously solved—and confirmed—the 3-D conformation of six of these unknown protein families.
“The similarity is impressive,” comments Johannes Söding of the Max Planck Institute for Biophysical Chemistry, who was not involved in the work. “And these aren’t simple, small proteins—most have several hundred amino acids and complex folds,” Söding says.
For more than two decades, researchers have been building computer models that predict the structure of a protein from its amino acid sequence, achieving only variable levels of success. Baker’s team’s method is based on an idea first floated 20 years ago.
Back then, Chris Sander, now at the Broad Institute of Massachusetts Institute of Technology and Harvard University, proposed that protein homology—the fact that many different organisms have proteins evolved from a common ancestor protein—could be used to make structure predictions. Homologous proteins have similar amino acid sequences, do similar jobs in cells, and typically have similar overall structures.
If the side chains of two amino acids in a protein are touching, they’ll want to continue touching throughout the course of evolution, Baker explains. “So if one happens to mutate so that it has a larger side chain, then the other amino acid must mutate so that it has a smaller side chain.”
Computer algorithms identify these correlated amino acids using sequence alignments of homologous proteins and advanced statistics. Because the correlated amino acids are typically close in space, this information can be used to predict 3-D structure.
To use this in silico strategy—a type of machine learning algorithm (see page 33)—Baker and coworkers needed many known protein sequences: The more sequences available to compare, the more reliable the modeling outcome. To boost the size of their data set, the researchers collected some of their “known” data from an unexpected source: microbial metagenomics research projects. In these projects, researchers directly sequence DNA from soil, ocean, and gut microbiomes. “No one ever conceived of using these data for protein structure predictions,” Baker says, but the metagenome projects helped the team improve the accuracy of its predictions.
“It’s a brute force, big-data strategy, and it seems to work better than anything else to date,” Söding notes.
This article has been translated into Spanish by Divulgame.org and can be found here.




Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter