Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Infectious disease
Covid-19
What do we know about the novel coronavirus’s 29 proteins?
These biomolecules could hold clues to why the virus is so infectious and to how to stop it
by Alla Katsnelson, special to C&EN
April 1, 2020
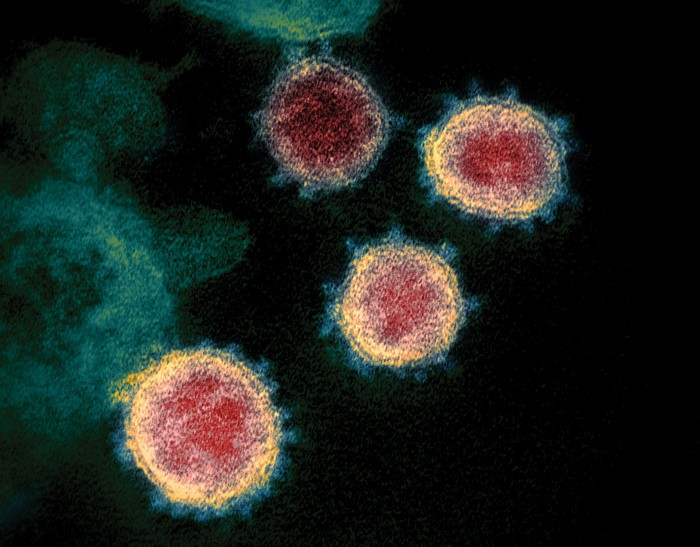
Scientists across the globe are gunning to understand the novel coronavirus, called SARS-CoV-2, and what makes it so contagious and deadly.
Several members of the coronavirus family infect humans: four cause the common cold and two—SARS-CoV and MERS-CoV—have triggered dangerous epidemics. The novel coronavirus’s closest kin is SARS-CoV, which jumped species from bats to civets to humans to cause the severe acute respiratory syndrome (SARS) epidemic of 2002–2003. That SARS outbreak infected over 8,000 people. The current coronavirus pandemic has infected more than 880,000 people and killed more than 44,000 as of April 1, according to data from Johns Hopkins University.
Support nonprofit science journalism
C&EN has made this story and all of its coverage of the coronavirus epidemic freely available during the outbreak to keep the public informed. To support us:
Donate Join Subscribe
“From a molecular perspective, figuring out why the virus is so much more transmissible than past viruses is where we should be looking right now,” says Robert Kirchdoerfer, a structural biologist at the University of Wisconsin–Madison who studies how coronaviruses fuse with host cells.
While people infected with the early 2000s SARS virus showed severe symptoms almost right away, people infected with SARS-CoV-2 can spread the virus before they show symptoms or when they are just mildly sick. A study published in Nature reports that patients shed virus most efficiently in the first week of illness, when their symptoms are mild (2020, DOI: 10.1038/s41586-020-2196-x). The US Centers for Disease Control and Prevention now estimates that 25% of people infected with the virus show no symptoms.
“That gives it an advantage to spread,” says Melanie Ott, a virologist at the University of California, San Francisco, and a member of the Quantitative Biosciences Institute’s COVID consortium. There must be something in the virus’s biological makeup that facilitates this silent transmission and that makes its effect on its human hosts so variable. But researchers don’t yet know what it is.
The RNA genome of SARS-CoV-2 has 29,811 nucleotides, encoding for 29 proteins, though one may not get expressed. Studying these different components of the virus, as well as how they interact with our cells, is already yielding some clues, but much remains to be learned, Ott says.
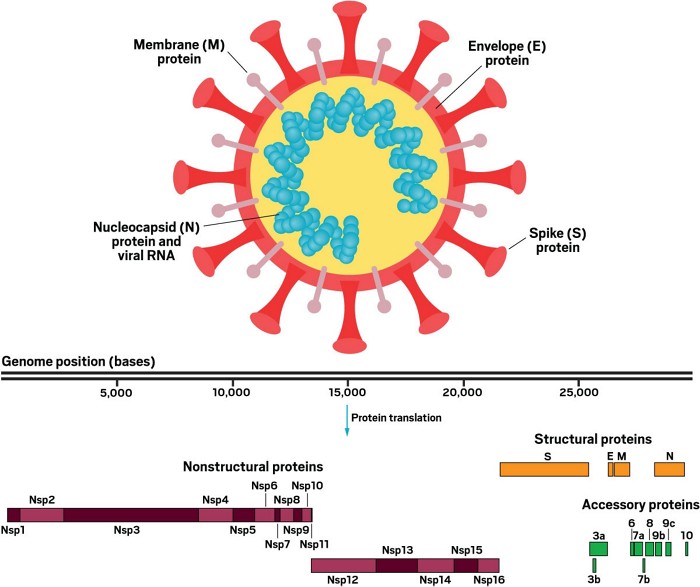
The spike
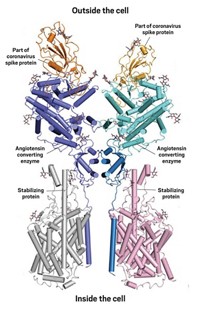
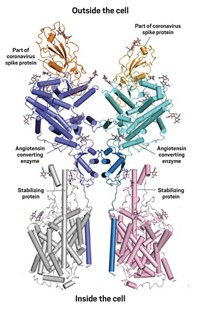
Coronaviruses are named for the crown of protein spikes covering their outer membrane surface. Early work on the novel coronavirus has focused on these spike proteins—also called S proteins—because they are the keys that the virus uses to enter host cells. In both SARS-CoV and SARS-CoV-2, the S protein binds to a receptor called angiotensin converting enzyme 2 (ACE2) to hack its way into host cells.
At the amino acid level, the spike proteins on SARS-CoV-2 are about 80% identical to those on SARS-CoV. “The residues which interact with ACE2 are conserved, compared to [the first] SARS, but the residues in between are different, and there are also some insertions,” says Rolf Hilgenfeld, a structural biologist who studies coronaviruses at the University of Lübeck.
Studies so far suggest that the new virus’s spike proteins bind to ACE2 significantly more strongly than those of SARS-CoV. “That is probably one of the reasons it spreads more easily and is more infectious,” Hilgenfeld says.
A non-peer-reviewed preprint posted on medRxiv on March 27 reports that ACE2 is expressed especially strongly in the lungs of people with lung diseases (2020, DOI: 10.1101/2020.03.21.20040261). That observation may partly explain how these underlying conditions make people more susceptible to infection by the novel coronavirus.
Stronger binding with ACE2 isn’t the only clue to the virus’s power. S proteins are made up of two segments, S1 and S2, that must be cleaved at two sites to expose a peptide that initiates fusion with ACE2 on a host cell. A mutation in the SARS-CoV-2 S protein allows an enzyme called furin, which is made by many types of human cells, to do the first cut. This ability to get human enzymes to do its work primes the virus for fusing with ACE2—providing another possible explanation for this virus’s infectiousness, says Carolyn Machamer, a cell biologist at Johns Hopkins University who studies the basic biology of coronaviruses. “It can get that first clip before it even comes into contact with a receptor.”
Researchers are working to identify the best way to interfere with the S protein’s interaction with ACE2. ACE2 is present in many organs throughout the body and interfering with it may have side effects, so researchers want to avoid hitting the receptor and are instead developing antibodies or peptides that bind and disable specific segments of the S protein.
The other 28
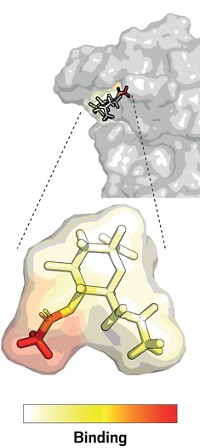
Of the 29 SARS-CoV-2 proteins, four make up the virus’s actual structure, including the S protein. One group of the other 25 coronavirus proteins regulates how the virus assembles copies of itself and how it sneaks past the host immune system. These so-called nonstructural proteins are expressed as two huge polyproteins that are then cleaved into 16 smaller proteins. An enzyme called the main protease, which performs 11 of those cleavages, is also a highly promising drug target. Hilgenfeld and his colleagues recently reported the structure of the main protease and identified an inhibitor that can block it.
Many of the virus’s nonstructural proteins are still poorly understood. “I think we are just going to have to go brute force and probably piece through the genome and study a lot of these proteins individually,” says Anthony Fehr, a biologist at the University of Kansas. These studies will help scientists both understand the underlying biochemistry that gives SARS-CoV-2 its nasty kick and identify other ways to fight the virus.
Fehr’s lab works on a nonstructural protein called NSP3, a component of which blocks the host’s efforts to fight off the virus. In particular, this protein shuts down host enzymes called PARPs, which prevent viruses from replicating, and interferes with cellular calls for the release of virus-fighting immune proteins called interferons. This protein could be a drug target, he says. “The virus would be dead in the water if it didn’t have a way to counter the interferon response.”
The third group of proteins in the novel coronavirus are accessory proteins. Coronaviruses don’t need these proteins to replicate in a test tube, but they do need the molecules to counteract the host’s innate immune system. However, accessory proteins are the least-well understood. “I’m interested in how the differences in these small proteins affect pathogenesis,” says Susan Weiss, a microbiologist who studies coronaviruses at the University of Pennsylvania. Her lab has studied accessory proteins in other coronaviruses; they plan to determine how mutations in these genes affect SARS-CoV-2’s ability to counteract host immune response and replicate.
Structural biology continues to be a key method for studying the new virus’s proteins. The differences between SARS-CoV-2 and the earlier SARS-CoV, “must have their basis in protein structure, but it’s quite difficult to correlate the 3-D structures—if you even have the 3-D structures—with differences in function,” Hilgenfeld says.
However, researchers have other tools to investigate SARS-CoV-2, says Susan Daniel, a biomolecular engineer at Cornell University who studies how coronaviruses enter cells. For example, electron spin resonance can provide information about how peptides involved in viral fusion get buried in the membranes of host cells; circular dichroism spectroscopy can provide insight about how alpha-helical structures within viral proteins change under specific conditions; isothermal titration calorimetry can hint at how viral peptides interact with various ions, which can give clues about their conformations; and nuclear magnetic resonance can provide important structural information not found in crystal structures.
Meanwhile, other labs are rushing to explore how differences in immune system responses might affect how severely ill an infected person may get. For example, Ott’s lab is beginning to infect organoids, which are models of lungs and other organs made of complex cell cultures, with the virus to watch how infections proceed. “The scientific community has come together in an unprecedented way,” Ott says. “Everybody basically has shifted their attention to the virus and is looking for ways to contribute meaningfully.”
CORRECTION
This story was updated on April 13, 2020, to correct a statistic about the 2002–03 SARS epidemic. That virus didn't kill more than 8,000 people worldwide; it infected that many. It killed over 700.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter