Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Synthetic Biology
Designer proteins prompt animals to make protective antibodies
The approach could help in the development of new vaccines and therapeutics
by Laura Howes
May 14, 2020
| A version of this story appeared in
Volume 98, Issue 19
Researchers at the Swiss Federal Institute of Technology, Lausanne (EPFL), report that they have used computational protein design to make small proteins that prod animals to develop a protective immune response against a specific virus (Science 2020, DOI: 10.1126/science.aay5051). Using this de novo process to design proteins for vaccines is a “lovely piece of work,” says Dek Woolfson at the University of Bristol, who was not involved in the research.
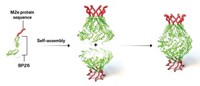
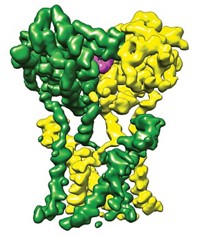
Most vaccines use inactivated bacteria or viruses or isolated parts of the infectious organism as targets for the immune system to develop antibodies against. Bruno Correia and his team wanted to make a vaccine against respiratory syncytial virus (RSV), which causes cold-like symptoms. They looked at a protein that RSV uses to infect cells and identified areas on the protein that antibodies target, called epitopes. The researchers then used computational design and screening to create functional protein mimics of the epitopes with the right shape and localized chemistry.
The team then created nanoparticles with three designed proteins on the outside and injected them into mice. After injection, the mice developed antibodies against RSV, although not as many as when the researchers injected the mice with the RSV protein. When they inoculated the mice first with the full RSV protein and then gave the mice booster shots of the designer protein particles, the antibody response refocused toward producing more neutralizing antibodies. The designer proteins also had the same effect in nonhuman primates.
Correia says his team doesn’t plan to use its process on the novel coronavirus SARS-CoV-2, adding that the vaccines currently in development will be the best way of fighting COVID-19. But, he says, this approach could help fight pathogens that are hard to develop vaccines for, or help top up people’s antibody response as they age.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter