Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Antibiotics
Phylogenetic screen finds antibiotics that work in a new way
Newly discovered compounds trap bacteria so they can’t grow or divide
by Laura Howes
February 16, 2020
| A version of this story appeared in
Volume 98, Issue 7

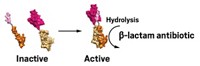
Antibiotic resistance is a tough problem—one some biologists worry cannot be solved with a business-as-usual approach to drug development. So researchers led by Gerard D. Wright at McMaster University tried out a new way of searching for antibiotic compounds. The team built a phylogenetic tree of gene clusters that code for glycopeptide antibiotics and then looked for self-resistance genes, which help bacteria protect themselves from the antibiotics that they make. The researchers found resistance genes for glycopeptide antibiotics on one large branch of the tree but not another, suggesting to the researchers that the antibiotics from the branch without the resistance genes might be part of a new functional family. So they searched that branch for antimicrobial compounds and purified two of them, including a newly found metabolite they named corbomycin. Corbomycin treats methicillin-resistant Staphylococcus aureus infections in mice, and it works in a novel way. The team tested the compounds and found that they prevent the breakdown of the peptidoglycan matrix in bacterial cell walls, effectively “caging” them so they cannot grow or divide (Nature 2020, DOI: 10.1038/s41586-020-1990-9).


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter