Advertisement
Grab your lab coat. Let's get started
Welcome!
Welcome!
Create an account below to get 6 C&EN articles per month, receive newsletters and more - all free.
It seems this is your first time logging in online. Please enter the following information to continue.
As an ACS member you automatically get access to this site. All we need is few more details to create your reading experience.
Not you? Sign in with a different account.
Not you? Sign in with a different account.
ERROR 1
ERROR 1
ERROR 2
ERROR 2
ERROR 2
ERROR 2
ERROR 2
Password and Confirm password must match.
If you have an ACS member number, please enter it here so we can link this account to your membership. (optional)
ERROR 2
ACS values your privacy. By submitting your information, you are gaining access to C&EN and subscribing to our weekly newsletter. We use the information you provide to make your reading experience better, and we will never sell your data to third party members.
Biochemistry
Lanthanide-binding proteins could unlock new sources of valuable elements
The discovery of bacteria that depend on lanthanides is delivering new ways to detect and extract crucial technology metals
by Mark Peplow, special to C&EN
February 7, 2022
| A version of this story appeared in
Volume 100, Issue 5

Credit: Yang H. Ku/C&EN/Shutterstock
In brief
The lanthanide elements were once thought to have no role in biology—until scientists discovered bacteria that cannot thrive without them. These microbes contain proteins and other molecules that capture, transport, and utilize lanthanides. Chemists are now using this biochemical ensemble, collectively known as the lanthanome, to extract lanthanides from waste materials so that the prized elements can be used in high-tech applications such as lasers and strong magnets. A lanthanome protein called lanmodulin can even purify actinides that are showing promise as medical radiotherapies. Meanwhile, biologists are discovering that lanthanide-loving bacteria are remarkably common and may have a major impact on the chemistry of their environments.
You can smell the Solfatara volcanic crater long before you see it on the drive west from Naples, Italy. The wide expanse of gray ash is pockmarked with fissures and fumaroles, some stained vivid yellow by sulfur as they huff clouds of steam laced with noxious gases. On one side, a pond of hot, gray mud bubbles and burps vigorously.
In 2005, Arjan Pol, a microbiologist at Radboud University, gingerly approached this mud pot, armed with some rather unconventional sampling equipment. “It was a little stainless steel beaker, which my wife used to heat the milk for cappuccino,” he recalls. “I fixed it with some tape to a broomstick. And that was it. We just scooped the water out of the mud pot, so I had to go relatively close to it.”
Pol was part of a team hoping to find microorganisms that could metabolize carbon disulfide in the mud’s hostile environment—a highly acidic pH 2 and a sweltering 60 °C. But his sample held an even more remarkable catch. Back in the lab, the researchers eventually realized that they had captured a bacterium that simply could not survive without a supply of lanthanides (Environ. Microbiol. 2014, DOI: 10.1111/1462-2920.12249).
Conventional wisdom presumed that the 15 lanthanide elements near the bottom of the periodic table had no role in biology because their ions generally prefer a +3 oxidation state that forms highly insoluble compounds in neutral water, limiting the elements’ bioavailability. But the acidic water in the mud pot was able to hold much higher concentrations of lanthanides, providing a rich broth to sustain the microbe that Pol and his colleagues named Methylacidiphilum fumariolicum SolV.

Over the past decade, it has become clear that lanthanide-using bacteria are far more widespread than anyone had ever imagined. Microbiologists have discovered many species that use a suite of specialized molecules to bind, transport, and employ these elements. This burgeoning band of biomolecules has been dubbed the lanthanome (ACS Cent. Sci. 2019, DOI: 10.1021/acscentsci.9b00642).
Chemists say members of the lanthanome could be adapted as sensors to identify new sources of lanthanides, which are in huge demand for applications as diverse as medical imaging, smartphones, and the generators in wind turbines. The lanthanome could also provide better ways to extract and recycle these metals, which together with scandium and yttrium are known as rare earth elements. “It’s absolutely incredible what has already been accomplished by adopting this natural machinery,” says Eric J. Schelter, a chemist at the University of Pennsylvania who studies the lanthanome. “These efficient, sustainable methods that give us access to new ores are potentially going to make a big impact on our ability to deliver materials for the clean energy transformation.”
Switched-on lanthanides
Methane provides the main source of energy for M. fumariolicum SolV, which oxidizes methane to methanol, then to formate, and ultimately to carbon dioxide. The Radboud University team, led by Huub J. M. Op den Camp, found that the methanol oxidation step relies on a methanol dehydrogenase (MDH) enzyme called XoxF, which carries a lanthanide ion at its heart. Without lanthanides—commonly abbreviated collectively as Ln—this crucial reaction sequence halts, and the microbe cannot grow.
M. fumariolicum was not the first bacterium known to host an Ln-MDH. In 2011 and 2012, a team led by Keiichi Kawai at Gifu University reported that several other bacteria carried this enzyme (J. Biosci. Bioeng. 2011, DOI: 10.1016/j.jbiosc.2010.12.017 and DOI: 10.1016/j.jbiosc.2011.01.015), although in the absence of lanthanides they could fall back on a more conventional calcium-dependent MDH called MxaF (PLOS One 2012, DOI: 10.1371/journal.pone.0050480). Add a dash of lanthanides, however, and these bacteria happily perform what is known as a lanthanide switch, reverting to Ln-MDH (J. Bacteriol. 2016, DOI: 10.1128/JB.00937-15).

“It’s remarkable that even nanomolar concentrations of lanthanides are enough to trigger the lanthanide pathway,” says bioinorganic chemist Lena J. Daumann of Ludwig Maximilian University Munich, who works on the lanthanome and collaborates with the Radboud team. The lanthanide switch most likely offers the bacteria a benefit, perhaps because Ln-MDH is better than Ca-MDH at catalyzing methanol oxidation.
XoxF and MxaF rely on a cofactor called pyrroloquinoline quinone (PQQ), which is embedded in their active sites. Work on model compounds by Schelter and others suggests that the oxidation mechanism involves a hydride transfer from methanol to PQQ (J. Am. Chem. Soc. 2017, DOI: 10.1021/jacs.7b12318). In the active site, a lanthanide ion (LnIII) acts as a Lewis acid that accepts an electron pair from PQQ to facilitate the reaction.
Schelter’s team is making additional model compounds to understand the differences between Ln-MDH and Ca-MDH, and Schelter hopes that this could eventually help create new methanol oxidation catalysts. “If we can take advantage of a highly optimized set of reactions from nature, it might be useful in a methanol fuel cell,” he says.
Researchers have so far identified at least five types of Ln-MDHs and several analogous ethanol dehydrogenases (J. Bacteriol. 2016, DOI: 10.1128/JB.00478-16), all of which have a strong preference for the lighter lanthanides. Daumann and her colleagues found that neodymium—the fourth element in the lanthanide series—produced the highest catalytic activity, probably because it offered the best combination of Lewis acidity and coordination abilities in the active site (Dalton Trans. 2018, DOI: 10.1039/c8dt01238e). She is now using these slight differences in PQQ-lanthanide binding to tease apart mixtures of lanthanides (Chem.—Eur. J. 2020, DOI: 10.1002/chem.202002653).
Getting a grip
Daumann’s work could feed into a much broader effort to sate the world’s hunger for lanthanides. In general, they are spread very thinly in Earth’s crust—there are few known concentrated reserves in ores. What’s more, the chemical similarity of lanthanide ions means that separating them from one another is exceedingly difficult.
Current industrial extraction methods typically rely on large amounts of acid, organic solvents, and binding agents to draw rare earth elements out of ores. Such extraction is costly and energy intensive and produces huge amounts of waste that can damage the environment if improperly handled. These difficulties also hamper lanthanide recycling from industrial scrap and electronic waste (Science 2019, DOI: 10.1126/science.aau7628).

Yet lanthanide-using bacteria are able to gather lanthanides from very dilute sources with much less fuss. Researchers think that certain lanthanome proteins could help make lanthanide extraction more efficient and reduce its environmental footprint, making it commercially viable to extract lanthanides from low-grade sources such as runoff from mines and electronic waste.
In 2018, Joseph A. Cotruvo Jr. and colleagues at Pennsylvania State University discovered a key part of the lanthanome that could help exploit these untapped sources: a small protein called lanmodulin, which has an astonishing affinity for lanthanides (J. Am. Chem. Soc. 2018, DOI: 10.1021/jacs.8b09842). Lanmodulin’s biological role is still uncertain, although it may be involved in the selective uptake and handling of lanthanides in microbes.
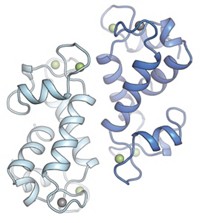
The protein wraps LnIII in a tight embrace using distinctive helix-loop-helix structures called EF hands, which are widely found in other biomolecules. When LnIII binds to the first of lanmodulin’s EF hands, it significantly changes the shape of the protein so that it is perfectly configured to bind a second LnIII—an effect called cooperative binding. A third EF hand can then more weakly bind yet another lanthanide ion (Biochemistry 2019, DOI: 10.1021/acs.biochem.8b01019).
Growing reputation

Cotruvo suggests that cooperative binding helps emphasize small differences in the ionic radius and coordination preference among metal ions, which makes lanmodulin incredibly good at selecting lanthanides from a soup of other ions. Mere picomolar concentrations of lighter lanthanides can induce lanmodulin’s conformational change, whereas it takes near-millimolar concentrations of calcium to have the same effect.
The protein is already showing promise in various applications. For example, Cotruvo’s team has transformed it into a sensor for terbium, a metal used in the phosphors of some energy-efficient lights. The team equipped lanmodulin’s EF hands with tryptophan residues that can absorb UV light and transfer some of the energy to terbium bound by the protein so that the ions glow green. The modified protein could detect nanomolar concentrations of TbIII in runoff from mines, even when other metal ions are present at concentrations 100,000 times as large (J. Am. Chem. Soc. 2021, DOI: 10.1021/jacs.1c06360). Researchers have estimated that acid drainage from mines in Pennsylvania and West Virginia alone could supply up to 3,400 metric tons (t) of rare earth elements per year (Min., Metall. Explor. 2019, DOI: 10.1007/s42461-019-0097-z), and Cotruvo suggests that the sensor might help identify the most fruitful locations for extraction.
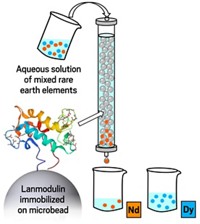
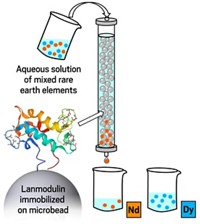
Cotruvo has also collaborated with Gauthier Deblonde at Lawrence Livermore National Laboratory to use lanmodulin to gather lanthanides from electronic waste and coal (Inorg. Chem. 2020, DOI: 10.1021/acs.inorgchem.0c01303). Anchored onto porous beads in a column, lanmodulin can separate lighter and heavier lanthanides from one another, even when they are vastly outnumbered by other ions such as aluminum and calcium (ACS Cent. Sci. 2021, DOI: 10.1021/acscentsci.1c00724). Schelter says that if these approaches “could be scaled up, we now have opportunities to go after sources of lanthanides that we’ve never had access to before.”

Lanmodulin is even more enthusiastic about actinides. Daumann found that it binds americium particularly well, for example (Chem. Sci. 2021, DOI: 10.1039/D1SC04827A), and Deblonde has used lanmodulin to separate radioactive americium from neptunium, a particularly challenging task in reprocessing spent nuclear fuel (J. Am. Chem. Soc. 2021, DOI: 10.1021/jacs.1c07103).
Lanmodulin could also help furnish the world with actinides for medical applications. Actinium-225, for example, decays by emitting an α particle and is being investigated for use in targeted cancer therapy. But the isotope is typically produced in particle accelerators, so supplies are limited and expensive.
Lanmodulin may provide an alternative source of a different actinium isotope for researching the chemistry of such therapeutic complexes. Deblonde’s team found that lanmodulin can isolate actinium-228 from readily available samples of its parent isotope, thorium-232, even in the presence of billions of times greater concentrations of other metal ions (Sci. Adv. 2021, DOI: 10.1126/sciadv.abk0273). “Rather than relying on the production cycle of big isotope producers, you can be autonomous and do your research at a much lower cost,” Deblonde says.
The ubiquitous lanthanome
While chemists explore applications for the lanthanome, biologists are making a slew of fresh discoveries about lanthanide-using bacteria. “Everywhere we look it seems more and more common to find bacteria using lanthanides,” says Jennifer B. Glass of the Georgia Institute of Technology, who has studied the distribution of lanmodulin genes. Various teams have found genetic traces of lanthanide-using bacteria in lakes, acidic peatlands, and even the soil of a park in England (FEMS Microbiol. Lett. 2020, DOI: 10.1093/femsle/fnaa165). “I think there are probably thousands of them out there,” Radboud’s Op den Camp says.

Lanmodulin and Ln-MDH reside in the periplasm of their host bacteria, a sort of cellular lobby that is sandwiched between a bacterium’s outer and inner cell membranes. But in 2020, a team led by N. Cecilia Martinez-Gomez at the University of California, Berkeley, reported that lanthanides were also being stored in polyphosphate granules in the cytoplasm of Methylorubrum extorquens AM1 (Sci. Rep. 2020, DOI: 10.1038/s41598-020-69401-4). These microbes have proteins in their inner and outer membranes that serve as gateways for lanthanides to enter the cell and access the cytoplasm (Mol. Microbiol. 2019, DOI:10.1111/mmi.14208). The same kinds of proteins are already known to act as portals for FeIII ions.
Advertisement
Like LnIII, FeIII is poorly soluble in water, so microbes often excrete chelating agents known as siderophores to bind the ion and carry it through the membrane. By analogy, some bacteria produce lanthanophores to harvest lanthanides in a similar way. “We are fairly confident that the excretion of these lanthanophores, at least for our microbes, is very important,” Martinez-Gomez says. She is now collaborating with Daumann to study the lanthanophores and exploit them to extract and separate lanthanides.
Martinez-Gomez also recently created a genetic variant of M. extorquens AM1 that is particularly good at acquiring gadolinium, commonly used in magnetic resonance imaging procedures (bioRxiv 2021, DOI: 10.1101/2021.06.12.448192). The preprint study has not been peer reviewed. People undergoing these procedures in the US alone collectively excrete about 20 t of gadolinium per year (Water Res. 2020, DOI: 10.1016/j.watres.2020.115966), and this waste can harm aquatic creatures. Martinez-Gomez hopes that her bacterium might offer a solution. “I think you can use this as a platform to clean medical waste or contaminated waters,” she says.
Meanwhile, researchers are also learning about the impact lanthanide-using bacteria have on their environment. After the 2010 Deepwater Horizon oil spill in the Gulf of Mexico, for instance, researchers found that naturally occurring lighter lanthanides had been removed by bacteria feasting in the methane-rich waters around the site (Sci. Rep. 2017, DOI: 10.1038/s41598-017-11060-z). Deblonde believes that the presence of lanmodulin or similar molecules could also complicate efforts to predict the behavior of nuclear waste at contaminated sites. Researchers modeling actinide transport in the environment generally regard elements like americium and curium as insoluble and immobile. “But if lanmodulin is around, they’re going to move, and that’s a concern,” Deblonde says.
This torrent of discoveries has turned lanthanide biochemistry into a fast-moving and competitive area. “A lot of people in the field are pretty young, so everyone is trying to make their mark,” Cotruvo says. It is also benefiting from interest at funding agencies such as the National Science Foundation, the US Department of Energy, and most recently the Defense Advanced Research Projects Agency, which in July issued a funding call for projects to develop scalable biobased separation and purification strategies for rare earth elements. “The timing of these discoveries is so perfect,” Cotruvo says. “We can leverage them to solve these really important scientific problems.”

Mark Peplow is a freelance science journalist based in Penrith, England. A version of this story appeared in ACS Central Science: cenm.ag/lanthanome.


Join the conversation
Contact the reporter
Submit a Letter to the Editor for publication
Engage with us on Twitter